RetrogeneDB ID: | retro_mmus_2769 | ||
Retrocopy location | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 6:99770637..99771121(+) | ||
| Located in intron of: | None | ||
Retrocopy information | Ensembl ID: | None | |
| Aliases: | None | ||
| Status: | NOVEL | ||
Parental gene information | Parental gene summary: | ||
| Parental gene symbol: | Btf3 | ||
| Ensembl ID: | ENSMUSG00000021660 | ||
| Aliases: | Btf3, 1700054E11Rik | ||
| Description: | basic transcription factor 3 [Source:MGI Symbol;Acc:MGI:1202875] |

Retrocopy-Parental alignment summary:
>retro_mmus_2769
AAGATGAAAGAAACGATCATGAACCAGGAAAAAACTTGCCAAACTGCAGGCACAAGTGCGCACTGGTGGGAAAAGAACTG
CTCGTAGAAAGAAGAAGGTTCACAGGCCAGCCACAGCAGAGGATAAGAAACTGCAGCTCTCCTTAGAGAAGTCAGGCGTA
AACAATATCTCTGGTATTGAAGAGGTGAGCATGTTTACAAACCAAAGAACAGGGATCCGTTTTAACAACCCTAAAGTTCA
GACATCCCTGGCAGCAGACACCCTCACCATTACAGGCCACGATGAGACAAAGCAGCGGACAGCAATGCTTTCCAGCATCC
TAAACCAGCTTGGTGCAGACGGTCTGACTAGTTTAAGGAGCCGGGCTGAAGCTCTGCCTAAACAATCCGTGGACGGAGAA
GCACCACTTACTCCTGGAGAGGAGGAGGAGGGTGAAGTTCCAGGACTGGTGGAGAATCTTGATGAGGCTTCTAAGAGCGA
GGCC
ORF - retro_mmus_2769 Open Reading Frame is not conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 79.14 % |
| Parental protein coverage: | 79.41 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected: | 1 |
Retrocopy - Parental Gene Alignment:
| Parental | QMKETIMNQEK-LAKLQAQVRIGGKGTARRKKKVVHRTATADDKKLQFSLKKLGVNNISGIEEVNMFTNQ |
| .MKETIMNQEK.LAKLQAQVR.GGK.TARRKKKV.HR.ATA.DKKLQ.SL.K.GVNNISGIEEV.MFTNQ | |
| Retrocopy | KMKETIMNQEK>LAKLQAQVRTGGKRTARRKKKV-HRPATAEDKKLQLSLEKSGVNNISGIEEVSMFTNQ |
| Parental | GTVIHFNNPKVQASLAANTFTITGHAETKQLTEMLPSILNQLGADSLTSLRRLAEALPKQSVDGKAPLAT |
| .T.I.FNNPKVQ.SLAA.T.TITGH.ETKQ.T.ML.SILNQLGAD.LTSLR..AEALPKQSVDG.APL.. | |
| Retrocopy | RTGIRFNNPKVQTSLAADTLTITGHDETKQRTAMLSSILNQLGADGLTSLRSRAEALPKQSVDGEAPLTP |
| Parental | GEDDDDEVPDLVENFDEASKNEA |
| GE....EVP.LVEN.DEASK.EA | |
| Retrocopy | GEEEEGEVPGLVENLDEASKSEA |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .05 RPM | 31 .21 RPM |
| SRP007412_cerebellum | 0 .00 RPM | 33 .00 RPM |
| SRP007412_heart | 0 .00 RPM | 33 .90 RPM |
| SRP007412_kidney | 0 .00 RPM | 50 .80 RPM |
| SRP007412_liver | 0 .00 RPM | 44 .76 RPM |
| SRP007412_testis | 0 .00 RPM | 108 .54 RPM |
RNA Polymerase II activity near the 5' end of retro_mmus_2769 was not detected
No EST(s) were mapped for retro_mmus_2769 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_126349 | 245 libraries | 217 libraries | 194 libraries | 309 libraries | 107 libraries |
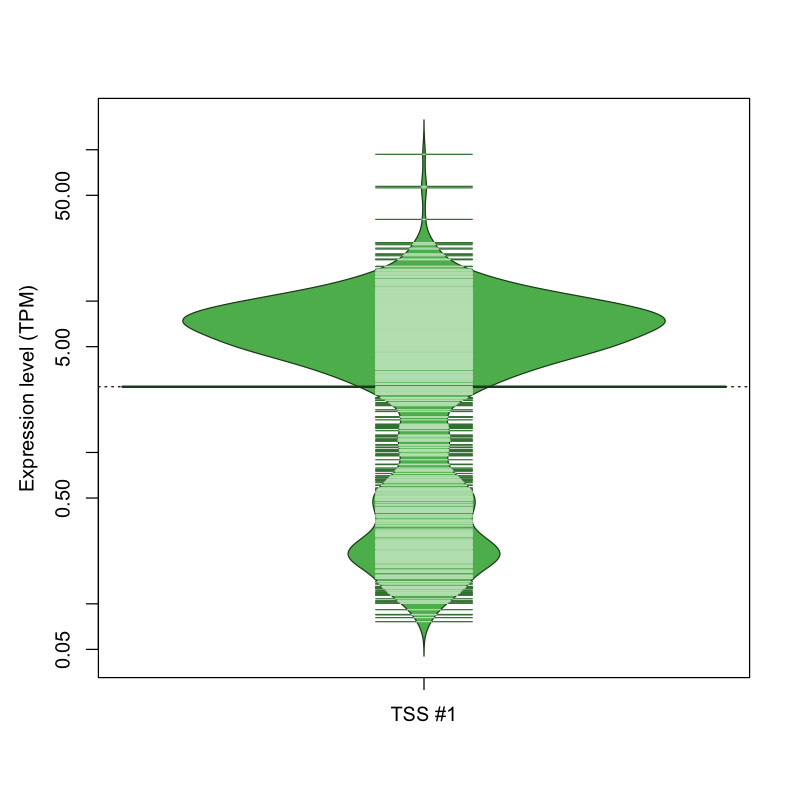
The graphical summary, for retro_mmus_2769 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
retro_mmus_2769 was not experimentally validated.