RetrogeneDB ID: | retro_mmus_185 | ||
Retrocopy location | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 6:67534752..67535706(-) | ||
| Located in intron of: | None | ||
Retrocopy information | Ensembl ID: | ENSMUSG00000051397 | |
| Aliases: | Tacstd2, C80403, EGP-1, GA733-1, Ly97, TROP2 | ||
| Status: | KNOWN_PROTEIN_CODING | ||
Parental gene information | Parental gene summary: | ||
| Parental gene symbol: | Epcam | ||
| Ensembl ID: | ENSMUSG00000045394 | ||
| Aliases: | Epcam, CD326, EGP, EGP-2, Egp314, Ep-CAM, EpCAM1, GA733-2, Ly74, TROP1, Tacsd1, Tacstd1, gp40 | ||
| Description: | epithelial cell adhesion molecule [Source:MGI Symbol;Acc:MGI:106653] |

Retrocopy-Parental alignment summary:
>retro_mmus_185
ATGGCGAGGGGCTTGGATCTAGCACCGCTGCTACTGCTACTGCTGGCGATGGCGACCCGCTTTTGCACGGCTCAGAGCAA
CTGTACATGCCCCACCAACAAGATGACGGTCTGCGACACAAATGGCCCAGGCGGGGTCTGCCAATGTCGGGCAATGGGCT
CACAGGTATTGGTCGACTGCTCCACGCTAACTTCCAAGTGCCTGCTGCTCAAGGCGCGCATGAGCGCCCGGAAGAGCGGC
CGCAGCCTGGTGATGCCGAGCGAGCACGCGATACTGGACAACGATGGCCTCTACGACCCGGAGTGTGACGACAAGGGCCG
CTTCAAGGCGCGCCAGTGCAACCAGACCTCGGTGTGCTGGTGCGTAAACTCGGTGGGCGTGCGCCGCACGGACAAGGGAG
ACCAAAGCCTGCGCTGCGACGAAGTGGTGCGAACCCACCACATCCTCATTGAGTTGCGCCACCGCCCGACCGACCGAGCC
TTCAACCACTCTGACCTAGACTCCGAGCTGCGGCGGCTCTTCCAAGAACGCTACAAGCTGCACCCCAGCTTCCTATCCGC
GGTACACTATGAGGAGCCCACCATTCAGATAGAGCTTCGGCAGAACGCGTCGCAGAAGGGCTTGAGAGACGTGGACATCG
CTGATGCCGCCTACTACTTCGAAAGGGACATTAAAGGCGAGTCACTGTTCATGGGCCGCCGCGGCCTGGACGTGCAGGTG
CGTGGGGAACCCCTGCATGTGGAGCGGACGCTCATCTACTACCTGGACGAGAAGCCCCCCCAGTTCTCCATGAAGCGCCT
CACCGCCGGCGTCATTGCCGTCATCGCTGTCGTCTCGGTAGCGGTAGTGGCTGGTGTGGTGGTCTTGGTGGTCACCAAAC
GGAGGAAGTCGGGCAAATACAAAAAGGTGGAGCTTAAGGAGCTGGGGGAGATGAGAAGCGAACCTAGCTTGTAG
ORF - retro_mmus_185 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 51.8 % |
| Parental protein coverage: | 95.87 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected: | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | LLLAVVTATLAAAQRDCVCDNYKLATSCSLNEYGECQCTSYGTQNTVICSKLASKCLAMKAEMTHSKSGR |
| LLL.........AQ..C.C...K..........G.CQC...G.Q..V.CS.L.SKCL..KA.M...KSGR | |
| Retrocopy | LLLLAMATRFCTAQSNCTCPTNKMTVCDTNGPGGVCQCRAMGSQVLVDCSTLTSKCLLLKARMSARKSGR |
| Parental | RI--KPEGAIQNNDGLYDPDCDEQGLFKAKQCNGTATCWCVNTAGVRRTDK-DTEITCSERVRTYWIIIE |
| ......E.AI..NDGLYDP.CD..G.FKA.QCN.T..CWCVN..GVRRTDK.D....C.E.VRT..I.IE | |
| Retrocopy | SLVMPSEHAILDNDGLYDPECDDKGRFKARQCNQTSVCWCVNSVGVRRTDKGDQSLRCDEVVRTHHILIE |
| Parental | LKHKERESPYDHQSLQTALQEAFTSRYKLNQKFIKNIMYENNVITIDLMQNSSQKTQDDVDIADVAYYFE |
| L.H........H..L...L...F..RYKL...F.....YE...I.I.L.QN.SQK...DVDIAD.AYYFE | |
| Retrocopy | LRHRPTDRAFNHSDLDSELRRLFQERYKLHPSFLSAVHYEEPTIQIELRQNASQKGLRDVDIADAAYYFE |
| Parental | KDVKGESLFHSSKSMDLRVNGEPLDLDPGQTLIYYVDEKAPEFSMQGLTAGIIAVIVVVSLAVIAGIVVL |
| .D.KGESLF......D..V.GEPL......TLIYY.DEK.P.FSM..LTAG.IAVI.VVS.AV.AG.VVL | |
| Retrocopy | RDIKGESLFMGRRGLDVQVRGEPLHVE--RTLIYYLDEKPPQFSMKRLTAGVIAVIAVVSVAVVAGVVVL |
| Parental | VISTRKKSAKYEKAEIKEMGEIHRE |
| V...R.KS.KY.K.E.KE.GE...E | |
| Retrocopy | VVTKRRKSGKYKKVELKELGEMRSE |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .07 RPM | 0 .14 RPM |
| SRP007412_cerebellum | 0 .00 RPM | 0 .09 RPM |
| SRP007412_heart | 0 .03 RPM | 0 .09 RPM |
| SRP007412_kidney | 17 .56 RPM | 70 .53 RPM |
| SRP007412_liver | 0 .03 RPM | 1 .69 RPM |
| SRP007412_testis | 0 .00 RPM | 24 .20 RPM |
RNA Polymerase II activity near the 5' end of retro_mmus_185 was not detected
No EST(s) were mapped for retro_mmus_185 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_124077 | 954 libraries | 63 libraries | 39 libraries | 16 libraries | 0 libraries |
| TSS #2 | TSS_124078 | 729 libraries | 98 libraries | 63 libraries | 46 libraries | 136 libraries |
| TSS #3 | TSS_124079 | 848 libraries | 121 libraries | 50 libraries | 49 libraries | 4 libraries |
The graphical summary, for retro_mmus_185 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
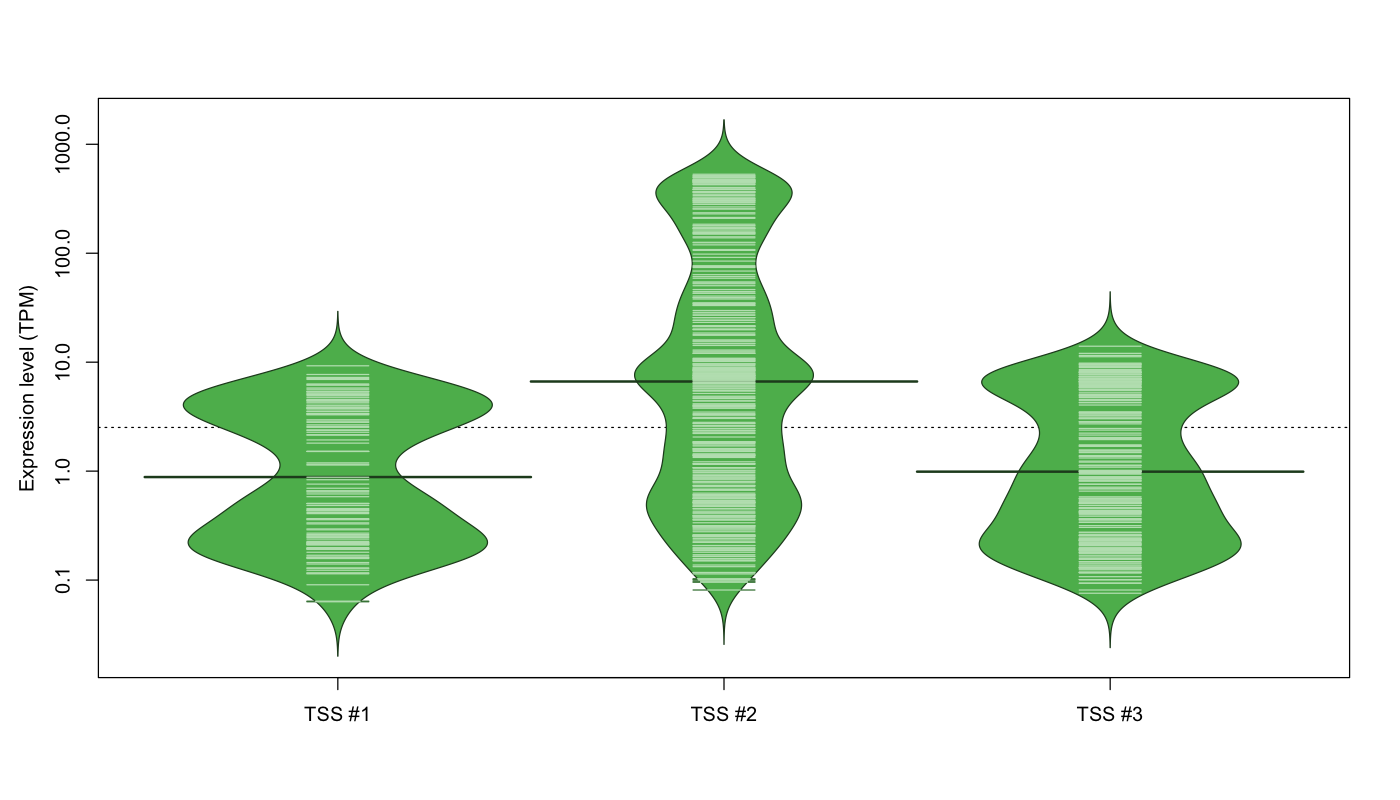
retro_mmus_185 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_185 has 1 orthologous retrocopies within eutheria group .| Species | RetrogeneDB ID |
|---|---|
| Rattus norvegicus | retro_rnor_252 |
Parental genes homology:
Parental genes homology involve 17 parental genes, and 19 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Bos taurus | ENSBTAG00000006474 | 1 retrocopy | |
| Choloepus hoffmanni | ENSCHOG00000013834 | 1 retrocopy | |
| Callithrix jacchus | ENSCJAG00000000624 | 1 retrocopy | |
| Dasypus novemcinctus | ENSDNOG00000011799 | 1 retrocopy | |
| Erinaceus europaeus | ENSEEUG00000009462 | 1 retrocopy | |
| Macropus eugenii | ENSMEUG00000015074 | 2 retrocopies | |
| Microcebus murinus | ENSMICG00000000151 | 1 retrocopy | |
| Macaca mulatta | ENSMMUG00000012390 | 1 retrocopy | |
| Monodelphis domestica | ENSMODG00000001077 | 1 retrocopy | |
| Mus musculus | ENSMUSG00000045394 | 1 retrocopy |
retro_mmus_185 ,
|
| Ochotona princeps | ENSOPRG00000004301 | 1 retrocopy | |
| Pelodiscus sinensis | ENSPSIG00000015680 | 1 retrocopy | |
| Pan troglodytes | ENSPTRG00000011902 | 1 retrocopy | |
| Rattus norvegicus | ENSRNOG00000015667 | 1 retrocopy | |
| Sus scrofa | ENSSSCG00000008429 | 1 retrocopy | |
| Ictidomys tridecemlineatus | ENSSTOG00000008631 | 1 retrocopy | |
| Tursiops truncatus | ENSTTRG00000004235 | 2 retrocopies |