RetrogeneDB ID: | retro_mmus_1131 | ||
Retrocopylocation | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 13:91261432..91262423(-) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | None | |
| Aliases: | None | ||
| Status: | NOVEL | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | Hnrnpk | ||
| Ensembl ID: | ENSMUSG00000021546 | ||
| Aliases: | Hnrnpk, Hnrpk, KBBP, NOVA | ||
| Description: | heterogeneous nuclear ribonucleoprotein K [Source:MGI Symbol;Acc:MGI:99894] |

Retrocopy-Parental alignment summary:
>retro_mmus_1131
CAGTACCAACATTATAAAGGAAGTGACTTTGATTGCGAGTTGAGACTGTTGATTCATCAGAGTCTGGCAGGAGGAATAAT
TGGTGTTAAAGGTGCTAAAATCAAAGAACTTCGAGAAAACACTCAGACAACAATCAAGCTTTTCCAGGAGTGCTGCCCTC
ACTCTACTGACAGAGTTGTTCTTATTGGAGGAAAACCTGATAGGGTTGTAGAATGCATTAAGATCATCCTTCACCTTATA
TCTGAGTCTCCTATCAAAGGACGTGCACAACCGTATGATCCCAACTTTTATGATGAGACCTATGATTATGGTGGTTTTAC
AATGATGTTTGATGACCGCTGAGGATGACCTGTGGGATTCCCCATGAGGGGAAGAGGTGGTTTTGACAGAATGCCTCCTG
GTCAGGGTGGGCGTCCTATGCCTCCTTCTAGAAGAGATTATGATGATATGAGCCCTTGTCGAGGACCTCCACCACCTCCT
CCTGGTCGAGGTGGCCGGGGTGGCAGCAGAGCCCGGAATCCTCTTCTTCCTCCACCACCACCCAGAGGGGGAGATCTAAT
GGCTTATGACAGAAGAGGAAGGCCTGGAGACCGCTATGATGGCATGGTTGGGTTCAGTGCTGATGAAACTTGGGACTCTG
CAATTGACACATGGAGCCCATCAGAATGGCAAATGGCTTATGAACCACAGGGTGGTTCTGGATATGACTATTCTTATGCA
GGGGGGTCGTGGCTCATATGGTGATCTTGGCGGACCTATTATCACTATACAAGTAACTATTCCCAAAGATTTGGCTGGAT
CTATTATTGCCAAAGGTGGTCAGCGGATTAAACAAATTTGTCATGAATCTGGAGCATCAATCAAAATTGATGAACCTTTA
GATGGATCTGAAGATCGGATCATTACCATTACAGGAACACAGGGCCAGATACAGAATACACAGTATTTGCTGCAGAACAG
TGTGAAGCAGTATGCAGATGTTGAAGGATTC
CAGTACCAACATTATAAAGGAAGTGACTTTGATTGCGAGTTGAGACTGTTGATTCATCAGAGTCTGGCAGGAGGAATAAT
TGGTGTTAAAGGTGCTAAAATCAAAGAACTTCGAGAAAACACTCAGACAACAATCAAGCTTTTCCAGGAGTGCTGCCCTC
ACTCTACTGACAGAGTTGTTCTTATTGGAGGAAAACCTGATAGGGTTGTAGAATGCATTAAGATCATCCTTCACCTTATA
TCTGAGTCTCCTATCAAAGGACGTGCACAACCGTATGATCCCAACTTTTATGATGAGACCTATGATTATGGTGGTTTTAC
AATGATGTTTGATGACCGCTGAGGATGACCTGTGGGATTCCCCATGAGGGGAAGAGGTGGTTTTGACAGAATGCCTCCTG
GTCAGGGTGGGCGTCCTATGCCTCCTTCTAGAAGAGATTATGATGATATGAGCCCTTGTCGAGGACCTCCACCACCTCCT
CCTGGTCGAGGTGGCCGGGGTGGCAGCAGAGCCCGGAATCCTCTTCTTCCTCCACCACCACCCAGAGGGGGAGATCTAAT
GGCTTATGACAGAAGAGGAAGGCCTGGAGACCGCTATGATGGCATGGTTGGGTTCAGTGCTGATGAAACTTGGGACTCTG
CAATTGACACATGGAGCCCATCAGAATGGCAAATGGCTTATGAACCACAGGGTGGTTCTGGATATGACTATTCTTATGCA
GGGGGGTCGTGGCTCATATGGTGATCTTGGCGGACCTATTATCACTATACAAGTAACTATTCCCAAAGATTTGGCTGGAT
CTATTATTGCCAAAGGTGGTCAGCGGATTAAACAAATTTGTCATGAATCTGGAGCATCAATCAAAATTGATGAACCTTTA
GATGGATCTGAAGATCGGATCATTACCATTACAGGAACACAGGGCCAGATACAGAATACACAGTATTTGCTGCAGAACAG
TGTGAAGCAGTATGCAGATGTTGAAGGATTC
ORF - retro_mmus_1131 Open Reading Frame is not conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 95.48 % |
| Parental protein coverage: | 75.23 % |
| Number of stop codons detected: | 2 |
| Number of frameshifts detected | 1 |
Retrocopy - Parental Gene Alignment:
| Parental | EYQHYKGSDFDCELRLLIHQSLAGGIIGVKGAKIKELRENTQTTIKLFQECCPHSTDRVVLIGGKPDRVV |
| .YQHYKGSDFDCELRLLIHQSLAGGIIGVKGAKIKELRENTQTTIKLFQECCPHSTDRVVLIGGKPDRVV | |
| Retrocopy | QYQHYKGSDFDCELRLLIHQSLAGGIIGVKGAKIKELRENTQTTIKLFQECCPHSTDRVVLIGGKPDRVV |
| Parental | ECIKIILDLISESPIKGRAQPYDPNFYDETYDYGGFTMMFDDRRGRPVGFPMRGRGGFDRMPPGRGGRPM |
| ECIKIIL.LISESPIKGRAQPYDPNFYDETYDYGGFTMMFDDR.G.PVGFPMRGRGGFDRMPPG.GGRPM | |
| Retrocopy | ECIKIILHLISESPIKGRAQPYDPNFYDETYDYGGFTMMFDDR*G*PVGFPMRGRGGFDRMPPGQGGRPM |
| Parental | PPSRRDYDDMSPRRGPPPPPPGRGGRGGSRARNLPLPPPPPPRGGDLMAYDRRGRPGDRYDGMVGFSADE |
| PPSRRDYDDMSP.RGPPPPPPGRGGRGGSRARN.PL.PPPPPRGGDLMAYDRRGRPGDRYDGMVGFSADE | |
| Retrocopy | PPSRRDYDDMSPCRGPPPPPPGRGGRGGSRARN-PLLPPPPPRGGDLMAYDRRGRPGDRYDGMVGFSADE |
| Parental | TWDSAIDTWSPSEWQMAYEPQGGSGYDYSYA-GGRGSYGDLGGPIITTQVTIPKDLAGSIIGKGGQRIKQ |
| TWDSAIDTWSPSEWQMAYEPQGGSGYDYSYA.GGRGSYGDLGGPIIT.QVTIPKDLAGSII.KGGQRIKQ | |
| Retrocopy | TWDSAIDTWSPSEWQMAYEPQGGSGYDYSYA>GGRGSYGDLGGPIITIQVTIPKDLAGSIIAKGGQRIKQ |
| Parental | IRHESGASIKIDEPLEGSEDRIITITGTQDQIQNAQYLLQNSVKQYADVEGF |
| I.HESGASIKIDEPL.GSEDRIITITGTQ.QIQN.QYLLQNSVKQYADVEGF | |
| Retrocopy | ICHESGASIKIDEPLDGSEDRIITITGTQGQIQNTQYLLQNSVKQYADVEGF |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 1 .78 RPM | 205 .71 RPM |
| SRP007412_cerebellum | 2 .82 RPM | 176 .42 RPM |
| SRP007412_heart | 2 .52 RPM | 146 .65 RPM |
| SRP007412_kidney | 3 .16 RPM | 198 .58 RPM |
| SRP007412_liver | 2 .62 RPM | 172 .59 RPM |
| SRP007412_testis | 2 .97 RPM | 166 .26 RPM |
RNA Polymerase II actvity near the 5' end of retro_mmus_1131 was not detected
No EST(s) were mapped for retro_mmus_1131 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_31483 | 99 libraries | 141 libraries | 581 libraries | 147 libraries | 104 libraries |
The graphical summary, for retro_mmus_1131 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
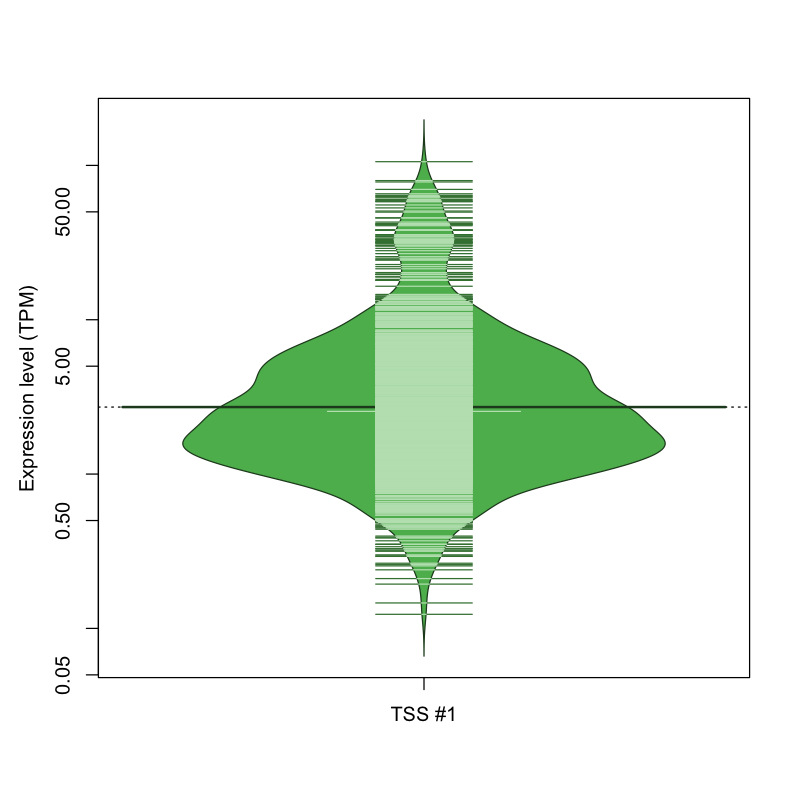
retro_mmus_1131 was not experimentally validated.