RetrogeneDB ID: | retro_mmus_3502 | ||
Retrocopylocation | Organism: | Mouse (Mus musculus) | |
| Coordinates: | X:105141134..105141940(+) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | ENSMUSG00000082245 | |
| Aliases: | None | ||
| Status: | KNOWN_PSEUDOGENE | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | Sms | ||
| Ensembl ID: | ENSMUSG00000071708 | ||
| Aliases: | Sms, AI427066, Gy, SPMSY, SpmST, gyro | ||
| Description: | spermine synthase [Source:MGI Symbol;Acc:MGI:109490] |

Retrocopy-Parental alignment summary:
>retro_mmus_3502
AAAGAATTGAGTCAAGACAGTACTGGGCGGGTGAAGCAATTACCACCCATAGTTCGAGGAGGGGCCATTGACAGATACTG
GCCTACTGCTGATGGGCGCCTGGCTGAGTATGACATAGATGAAGTGGTGTATGATGAAGATTCACCCTATCAGAACATTA
AAATCCTACACTCAGAGCAGTTTGGGAATACTCTCATCCTCAGCGGGGATGTTAATTTGGCAGAGAGTGACTTGGCATAT
ACCCGTGCCATCATGGGCAGTGGTAAAAAAGACTACACTGGCAAAGATGTACTGATTCTGGGAGGCGGCGATGGAGGCAT
ATTGTGTGAAATTGTCAAACTGAAGCCAAAGATGGTCACTATGGTAGAGATTGACCAAATGGTCACTGATGGATGTAAGA
AATGCATGCGGAGAACATGTGGAGATGTCTTAGACAATCTTAAAGGAGACTGCTATCAGGGTCTTATAGAAGACTGAATC
CCAGTACTGAAGAGGTTTGCCAAAGAAGGGAGAGAGTTTGATTATGTGATTAGTGATTTGACAGTTGTCCCTAACTCTAC
TCTCCAGAAGAAGATTGCACATGGGAGTTCCTCAGACTGATTCTCGACCTGTTGATGAAGGTTCTGAAACAGGCTGGAAA
ATACTTCACACAGGGGAACTGTGTCAACTTTACTGAAGCCCTGTCGCTCTATGACGAACAGCTGGGGTGTCTGTATTGTC
TTGTGGAATTCTCAAAAGAGATTGTCTGTGTCCCTTCATACTTGGAATTGTGGGTATTTTACACTGTTTGGAAGAAAGCT
GAGCCT
AAAGAATTGAGTCAAGACAGTACTGGGCGGGTGAAGCAATTACCACCCATAGTTCGAGGAGGGGCCATTGACAGATACTG
GCCTACTGCTGATGGGCGCCTGGCTGAGTATGACATAGATGAAGTGGTGTATGATGAAGATTCACCCTATCAGAACATTA
AAATCCTACACTCAGAGCAGTTTGGGAATACTCTCATCCTCAGCGGGGATGTTAATTTGGCAGAGAGTGACTTGGCATAT
ACCCGTGCCATCATGGGCAGTGGTAAAAAAGACTACACTGGCAAAGATGTACTGATTCTGGGAGGCGGCGATGGAGGCAT
ATTGTGTGAAATTGTCAAACTGAAGCCAAAGATGGTCACTATGGTAGAGATTGACCAAATGGTCACTGATGGATGTAAGA
AATGCATGCGGAGAACATGTGGAGATGTCTTAGACAATCTTAAAGGAGACTGCTATCAGGGTCTTATAGAAGACTGAATC
CCAGTACTGAAGAGGTTTGCCAAAGAAGGGAGAGAGTTTGATTATGTGATTAGTGATTTGACAGTTGTCCCTAACTCTAC
TCTCCAGAAGAAGATTGCACATGGGAGTTCCTCAGACTGATTCTCGACCTGTTGATGAAGGTTCTGAAACAGGCTGGAAA
ATACTTCACACAGGGGAACTGTGTCAACTTTACTGAAGCCCTGTCGCTCTATGACGAACAGCTGGGGTGTCTGTATTGTC
TTGTGGAATTCTCAAAAGAGATTGTCTGTGTCCCTTCATACTTGGAATTGTGGGTATTTTACACTGTTTGGAAGAAAGCT
GAGCCT
ORF - retro_mmus_3502 Open Reading Frame is not conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 90.74 % |
| Parental protein coverage: | 73.5 % |
| Number of stop codons detected: | 1 |
| Number of frameshifts detected | 1 |
Retrocopy - Parental Gene Alignment:
| Parental | KELSQDSTGRVKRLPPIVRGGAIDRYWPTADGRLVEYDIDEVVYDEDSPYQNIKILHSKQFGNILILSGD |
| KELSQDSTGRVK.LPPIVRGGAIDRYWPTADGRL.EYDIDEVVYDEDSPYQNIKILHS.QFGN.LILSGD | |
| Retrocopy | KELSQDSTGRVKQLPPIVRGGAIDRYWPTADGRLAEYDIDEVVYDEDSPYQNIKILHSEQFGNTLILSGD |
| Parental | VNLAESDLAYTRAIMGSGKEDYTGKDVLILGGGDGGILCEIVKLKPKMVTMVEIDQMVIDGCKKYMRRTC |
| VNLAESDLAYTRAIMGSGK.DYTGKDVLILGGGDGGILCEIVKLKPKMVTMVEIDQMV.DGCKK.MRRTC | |
| Retrocopy | VNLAESDLAYTRAIMGSGKKDYTGKDVLILGGGDGGILCEIVKLKPKMVTMVEIDQMVTDGCKKCMRRTC |
| Parental | GDVLDNLRGDCYQVLIEDCIPVLKMYAKEGREFDYVINDLTAVPIST-SPEEDSTWDFLRLILDLSMKVL |
| GDVLDNL.GDCYQ.LIED.IPVLK..AKEGREFDYVI.DLT.VP.ST.SPEED.TW.FLRLILDL.MKVL | |
| Retrocopy | GDVLDNLKGDCYQGLIED*IPVLKRFAKEGREFDYVISDLTVVPNST<SPEEDCTWEFLRLILDLLMKVL |
| Parental | KQDGKYFTQGNCVNLTEALSLYEEQLGRLYCPVEFSKEIVCVPSYLELWVFYTVWKKAKP |
| KQ.GKYFTQGNCVN.TEALSLY.EQLG.LYC.VEFSKEIVCVPSYLELWVFYTVWKKA.P | |
| Retrocopy | KQAGKYFTQGNCVNFTEALSLYDEQLGCLYCLVEFSKEIVCVPSYLELWVFYTVWKKAEP |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .02 RPM | 66 .02 RPM |
| SRP007412_cerebellum | 0 .00 RPM | 67 .91 RPM |
| SRP007412_heart | 0 .03 RPM | 24 .97 RPM |
| SRP007412_kidney | 0 .02 RPM | 31 .51 RPM |
| SRP007412_liver | 0 .00 RPM | 2 .79 RPM |
| SRP007412_testis | 0 .00 RPM | 2 .24 RPM |
RNA Polymerase II actvity near the 5' end of retro_mmus_3502 was not detected
No EST(s) were mapped for retro_mmus_3502 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_154752 | 369 libraries | 467 libraries | 229 libraries | 7 libraries | 0 libraries |
The graphical summary, for retro_mmus_3502 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
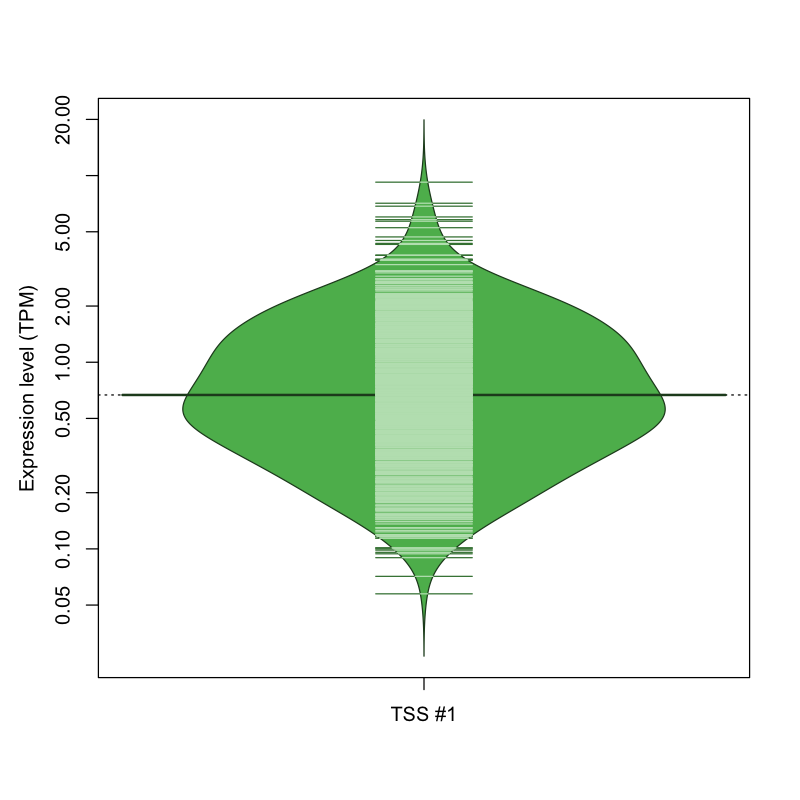
retro_mmus_3502 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_3502 has 0 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 14 parental genes, and 41 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Canis familiaris | ENSCAFG00000013284 | 3 retrocopies | |
| Callithrix jacchus | ENSCJAG00000033933 | 1 retrocopy | |
| Cavia porcellus | ENSCPOG00000006176 | 2 retrocopies | |
| Echinops telfairi | ENSETEG00000010098 | 2 retrocopies | |
| Homo sapiens | ENSG00000102172 | 5 retrocopies | |
| Gorilla gorilla | ENSGGOG00000003123 | 3 retrocopies | |
| Myotis lucifugus | ENSMLUG00000013185 | 3 retrocopies | |
| Macaca mulatta | ENSMMUG00000020392 | 2 retrocopies | |
| Monodelphis domestica | ENSMODG00000007897 | 1 retrocopy | |
| Mus musculus | ENSMUSG00000071708 | 6 retrocopies |
retro_mmus_2162, retro_mmus_2170, retro_mmus_2837, retro_mmus_3502 , retro_mmus_3628, retro_mmus_994,
|
| Nomascus leucogenys | ENSNLEG00000009955 | 4 retrocopies | |
| Pongo abelii | ENSPPYG00000020182 | 4 retrocopies | |
| Rattus norvegicus | ENSRNOG00000007688 | 2 retrocopies | |
| Tupaia belangeri | ENSTBEG00000012109 | 3 retrocopies |