RetrogeneDB ID: | retro_hsap_92 | ||
Retrocopylocation | Organism: | Human (Homo sapiens) | |
| Coordinates: | 8:88885011..88886199(-) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | ENSG00000176566 | |
| Aliases: | DCAF4L2, WDR21C | ||
| Status: | KNOWN_PROTEIN_CODING | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | DCAF4 | ||
| Ensembl ID: | ENSG00000119599 | ||
| Aliases: | DCAF4, WDR21, WDR21A | ||
| Description: | DDB1 and CUL4 associated factor 4 [Source:HGNC Symbol;Acc:20229] |

Retrocopy-Parental alignment summary:
>retro_hsap_92
ATGGAGAGCAAAAGACCGCGACTGCTCGAGGAAGCAGACAAGCAGAAAAAGACAGTCAGAGTGGGACTCAATGCACCTTC
CATGCTACGAAAGAACCAGCTAGGTTTCCTCAGATTCGCCAACTATTGCCGTATAGCTCGCGAGCTGCGTGTAAGCTGCA
TGCAGAGGAAAAAGGTCCAGATTCATAGCTGGGATCCCTCCTCTTTGGCAAGCGACCGATTTAACCGCATACTGGCGAAT
ACCAACACTGACCAGCTCTTCACAGTGAACCAAGTCGAAGCTGGAGGCTCCAAGTACGGCATCATCACCATGCGAGGCCT
GACGACCCCTGAGCTCCGGGTATACCCGCACAAAACCCTCTACGTCCCTAATCGGAAGGTGAATTCTATGTGCTGGGCCT
CACTGAATCACTTGGATTCCCACCTTCTGCTGTGCTTCGTGGGACTTGCAGATACTCCAAGCTGTGCCGTGCTGCTCCCA
GCGTCGCTGTTCATAGGTAGCTTCCCAGGAATGCGTCGGCCTGGCATGCTTTGCAGTTTCCAGATCCCTGATGCCTGGTC
CTGTGCCTGGTCCCTGAGCATCCACGCGTATCACTCTTTCAGTACAGGCTTGTCTCAGCAGGTCCTGTTGACCAACGTGG
TGACGGGACACCAGCAGTCATTTGGGACTAGCAGTGATGTCTTGGCCCAGCAGTTTGCAATCATGACTCCTTTGCTGTTT
AATGGCTGTCGCTCTGGGGAGATCTTTGGCATTGATCTGCGCTGTGGAAATCAAGGCAGCGGGTGGAAGGCCATTTGCCT
GTCCCATGATTCAGCAGTGACTTCTCTGCAAATCCTCCAAGATGGCCAATTCCTGGTGTCATCAGACATGACTGGAACTA
TCAAGCTGTGGGACTTGAGGGCCACTAAATGTGTAACACAGTACGAAGGTCATGTGAATAACTCCGCCTACCTACCCGTG
CATGTGAACGAAGAAGAAGGAGTCGTGGCGGCCGTGGGCCAGGACTGCTACACGAGAATCTGGAGCCTCCGTCATGGCCA
CCTGCTCACAACCATACCCTCCCCATACCCCGCCTCGGAGAACGACATTCCCAGTGTGGCCTTCTCTTCTCGCCTCGGGG
GCTTCCGAGGAGCACCAGGGCTGCTCATGGCTGTCCGGGAGGACCTTTATTGTTTCTCCTACGGTTAA
ATGGAGAGCAAAAGACCGCGACTGCTCGAGGAAGCAGACAAGCAGAAAAAGACAGTCAGAGTGGGACTCAATGCACCTTC
CATGCTACGAAAGAACCAGCTAGGTTTCCTCAGATTCGCCAACTATTGCCGTATAGCTCGCGAGCTGCGTGTAAGCTGCA
TGCAGAGGAAAAAGGTCCAGATTCATAGCTGGGATCCCTCCTCTTTGGCAAGCGACCGATTTAACCGCATACTGGCGAAT
ACCAACACTGACCAGCTCTTCACAGTGAACCAAGTCGAAGCTGGAGGCTCCAAGTACGGCATCATCACCATGCGAGGCCT
GACGACCCCTGAGCTCCGGGTATACCCGCACAAAACCCTCTACGTCCCTAATCGGAAGGTGAATTCTATGTGCTGGGCCT
CACTGAATCACTTGGATTCCCACCTTCTGCTGTGCTTCGTGGGACTTGCAGATACTCCAAGCTGTGCCGTGCTGCTCCCA
GCGTCGCTGTTCATAGGTAGCTTCCCAGGAATGCGTCGGCCTGGCATGCTTTGCAGTTTCCAGATCCCTGATGCCTGGTC
CTGTGCCTGGTCCCTGAGCATCCACGCGTATCACTCTTTCAGTACAGGCTTGTCTCAGCAGGTCCTGTTGACCAACGTGG
TGACGGGACACCAGCAGTCATTTGGGACTAGCAGTGATGTCTTGGCCCAGCAGTTTGCAATCATGACTCCTTTGCTGTTT
AATGGCTGTCGCTCTGGGGAGATCTTTGGCATTGATCTGCGCTGTGGAAATCAAGGCAGCGGGTGGAAGGCCATTTGCCT
GTCCCATGATTCAGCAGTGACTTCTCTGCAAATCCTCCAAGATGGCCAATTCCTGGTGTCATCAGACATGACTGGAACTA
TCAAGCTGTGGGACTTGAGGGCCACTAAATGTGTAACACAGTACGAAGGTCATGTGAATAACTCCGCCTACCTACCCGTG
CATGTGAACGAAGAAGAAGGAGTCGTGGCGGCCGTGGGCCAGGACTGCTACACGAGAATCTGGAGCCTCCGTCATGGCCA
CCTGCTCACAACCATACCCTCCCCATACCCCGCCTCGGAGAACGACATTCCCAGTGTGGCCTTCTCTTCTCGCCTCGGGG
GCTTCCGAGGAGCACCAGGGCTGCTCATGGCTGTCCGGGAGGACCTTTATTGTTTCTCCTACGGTTAA
ORF - retro_hsap_92 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 74.62 % |
| Parental protein coverage: | 79.6 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | MESKRLRLLQEEDRRKKIARMGFNASSMLRKSQLGFLNVTNYCHLAHELRLSCMERKKVQIRSMDPSALA |
| MESKR.RLL.E.D..KK..R.G.NA.SMLRK.QLGFL...NYC..A.ELR.SCM.RKKVQI.S.DPS.LA | |
| Retrocopy | MESKRPRLLEEADKQKKTVRVGLNAPSMLRKNQLGFLRFANYCRIARELRVSCMQRKKVQIHSWDPSSLA |
| Parental | SDRFNLILADTNSDRLFTVNDVKVGGSKYGIINLQSLKTPTLKVFMHENLYFTNRKVNSVCWASLNHLDS |
| SDRFN.ILA.TN.D.LFTVN.V..GGSKYGII....L.TP.L.V..H..LY..NRKVNS.CWASLNHLDS | |
| Retrocopy | SDRFNRILANTNTDQLFTVNQVEAGGSKYGIITMRGLTTPELRVYPHKTLYVPNRKVNSMCWASLNHLDS |
| Parental | HILLCLMGLAETPGCATLLPASLFVNSHPGIDRPGMLCSFRIPGAWSCAWSLNIQANNCFSTGLSRRVLL |
| H.LLC..GLA.TP.CA.LLPASLF..S.PG..RPGMLCSF.IP.AWSCAWSL.I.A...FSTGLS..VLL | |
| Retrocopy | HLLLCFVGLADTPSCAVLLPASLFIGSFPGMRRPGMLCSFQIPDAWSCAWSLSIHAYHSFSTGLSQQVLL |
| Parental | TNVVTGHRQSFGTNSDVLAQQFALMAPLLFNGCRSGEIFAIDLRCGNQGKGWKATRLFHDSAVTSVRILQ |
| TNVVTGH.QSFGT.SDVLAQQFA.M.PLLFNGCRSGEIF.IDLRCGNQG.GWKA..L.HDSAVTS..ILQ | |
| Retrocopy | TNVVTGHQQSFGTSSDVLAQQFAIMTPLLFNGCRSGEIFGIDLRCGNQGSGWKAICLSHDSAVTSLQILQ |
| Parental | DEQYLMASDMAGKIKLWDLRTTKCVRQYEGHVNEYAYLPLHVHEEEGILVAVGQDCYTRIWSLHDARLLR |
| D.Q.L..SDM.G.IKLWDLR.TKCV.QYEGHVN..AYLP.HV.EEEG...AVGQDCYTRIWSL....LL. | |
| Retrocopy | DGQFLVSSDMTGTIKLWDLRATKCVTQYEGHVNNSAYLPVHVNEEEGVVAAVGQDCYTRIWSLRHGHLLT |
| Parental | TIPSPYPASKADIPSVAFSSRLGGSRGAPGLLMAVGQDLYCYSY |
| TIPSPYPAS..DIPSVAFSSRLGG.RGAPGLLMAV..DLYC.SY | |
| Retrocopy | TIPSPYPASENDIPSVAFSSRLGGFRGAPGLLMAVREDLYCFSY |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| bodymap2_adipose | 0 .00 RPM | 6 .06 RPM |
| bodymap2_adrenal | 0 .00 RPM | 16 .95 RPM |
| bodymap2_brain | 0 .00 RPM | 7 .81 RPM |
| bodymap2_breast | 0 .00 RPM | 9 .14 RPM |
| bodymap2_colon | 0 .00 RPM | 4 .66 RPM |
| bodymap2_heart | 0 .00 RPM | 2 .27 RPM |
| bodymap2_kidney | 0 .00 RPM | 9 .27 RPM |
| bodymap2_liver | 0 .00 RPM | 4 .06 RPM |
| bodymap2_lung | 0 .00 RPM | 4 .83 RPM |
| bodymap2_lymph_node | 0 .00 RPM | 5 .57 RPM |
| bodymap2_ovary | 0 .00 RPM | 14 .11 RPM |
| bodymap2_prostate | 0 .00 RPM | 8 .56 RPM |
| bodymap2_skeletal_muscle | 0 .00 RPM | 7 .51 RPM |
| bodymap2_testis | 9 .65 RPM | 15 .52 RPM |
| bodymap2_thyroid | 0 .00 RPM | 21 .43 RPM |
| bodymap2_white_blood_cells | 0 .00 RPM | 5 .06 RPM |
RNA Polymerase II actvity may be related with retro_hsap_92 in 3 libraries
| ENCODE library ID | Target | ChIP-Seq Peak coordinates |
|---|---|---|
| ENCFF002CMI | POLR2A | 8:88886109..88886370 |
| ENCFF002CXN | POLR2A | 8:88885981..88886477 |
| ENCFF002CXO | POLR2A | 8:88886025..88886449 |
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_186975 | 1801 libraries | 12 libraries | 13 libraries | 2 libraries | 1 library |
| TSS #2 | TSS_186976 | 1789 libraries | 14 libraries | 19 libraries | 3 libraries | 4 libraries |
| TSS #3 | TSS_186977 | 1812 libraries | 6 libraries | 10 libraries | 1 library | 0 libraries |
The graphical summary, for retro_hsap_92 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1829 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
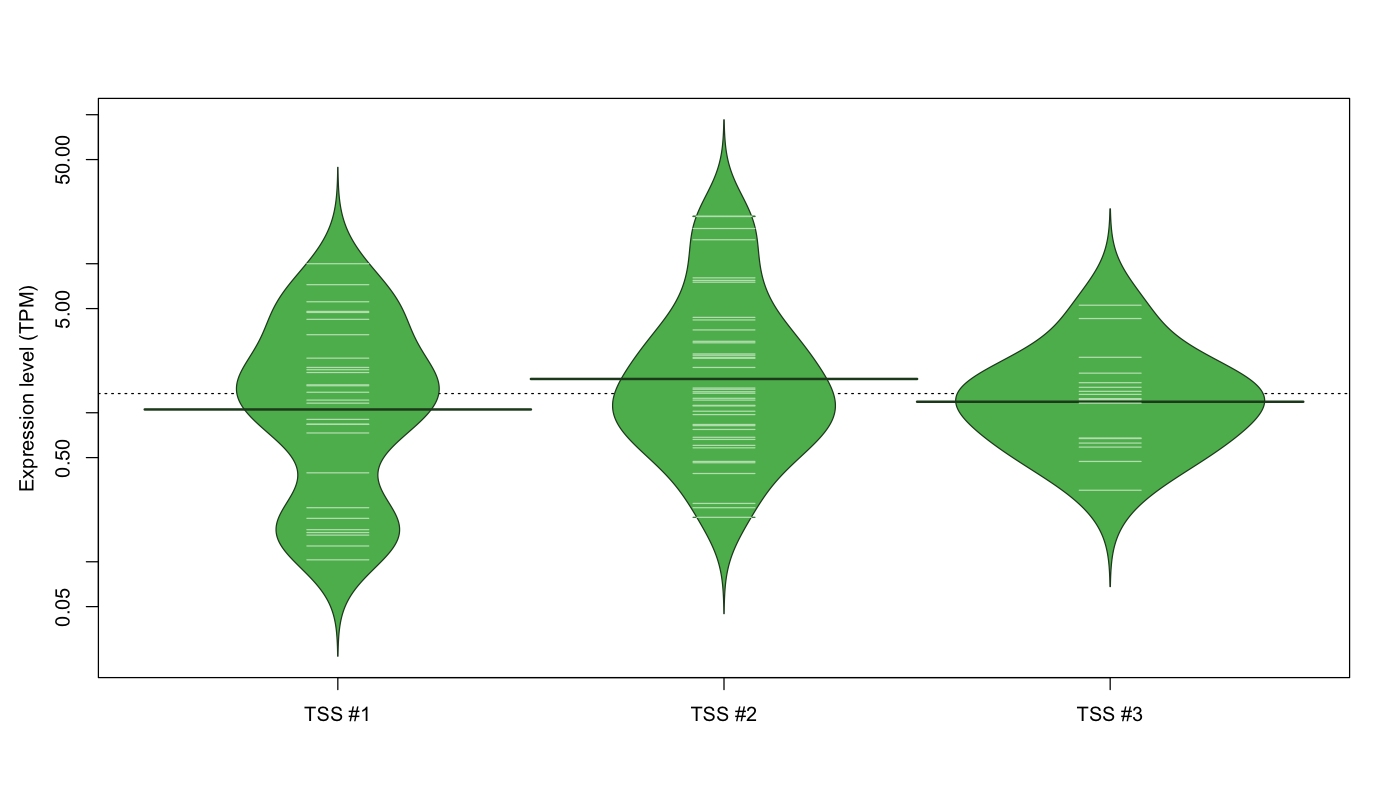
retro_hsap_92 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_hsap_92 has 3 orthologous retrocopies within eutheria group .| Species | RetrogeneDB ID |
|---|---|
| Pan troglodytes | retro_ptro_2774 |
| Gorilla gorilla | retro_ggor_2743 |
| Macaca mulatta | retro_mmul_264 |
Parental genes homology:
Parental genes homology involve 6 parental genes, and 11 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Cavia porcellus | ENSCPOG00000013878 | 1 retrocopy | |
| Homo sapiens | ENSG00000119599 | 2 retrocopies |
retro_hsap_101, retro_hsap_92 ,
|
| Gorilla gorilla | ENSGGOG00000003815 | 2 retrocopies | |
| Macaca mulatta | ENSMMUG00000003625 | 2 retrocopies | |
| Nomascus leucogenys | ENSNLEG00000002449 | 2 retrocopies | |
| Pan troglodytes | ENSPTRG00000006507 | 2 retrocopies |
Expression level across human populations :
| Library | Retrogene expression |
|---|---|
| CEU_NA11831 | 0 .00 RPM |
| CEU_NA11843 | 0 .00 RPM |
| CEU_NA11930 | 0 .00 RPM |
| CEU_NA12004 | 0 .00 RPM |
| CEU_NA12400 | 0 .00 RPM |
| CEU_NA12751 | 0 .00 RPM |
| CEU_NA12760 | 0 .00 RPM |
| CEU_NA12827 | 0 .00 RPM |
| CEU_NA12872 | 0 .00 RPM |
| CEU_NA12873 | 0 .00 RPM |
| FIN_HG00183 | 0 .00 RPM |
| FIN_HG00277 | 0 .00 RPM |
| FIN_HG00315 | 0 .00 RPM |
| FIN_HG00321 | 0 .00 RPM |
| FIN_HG00328 | 0 .00 RPM |
| FIN_HG00338 | 0 .00 RPM |
| FIN_HG00349 | 0 .00 RPM |
| FIN_HG00375 | 0 .00 RPM |
| FIN_HG00377 | 0 .00 RPM |
| FIN_HG00378 | 0 .00 RPM |
| GBR_HG00099 | 0 .00 RPM |
| GBR_HG00111 | 0 .00 RPM |
| GBR_HG00114 | 0 .00 RPM |
| GBR_HG00119 | 0 .00 RPM |
| GBR_HG00131 | 0 .00 RPM |
| GBR_HG00133 | 0 .00 RPM |
| GBR_HG00134 | 0 .00 RPM |
| GBR_HG00137 | 0 .00 RPM |
| GBR_HG00142 | 0 .00 RPM |
| GBR_HG00143 | 0 .00 RPM |
| TSI_NA20512 | 0 .00 RPM |
| TSI_NA20513 | 0 .00 RPM |
| TSI_NA20518 | 0 .00 RPM |
| TSI_NA20532 | 0 .00 RPM |
| TSI_NA20538 | 0 .00 RPM |
| TSI_NA20756 | 0 .00 RPM |
| TSI_NA20765 | 0 .00 RPM |
| TSI_NA20771 | 0 .00 RPM |
| TSI_NA20786 | 0 .00 RPM |
| TSI_NA20798 | 0 .00 RPM |
| YRI_NA18870 | 0 .00 RPM |
| YRI_NA18907 | 0 .00 RPM |
| YRI_NA18916 | 0 .00 RPM |
| YRI_NA19093 | 0 .00 RPM |
| YRI_NA19099 | 0 .00 RPM |
| YRI_NA19114 | 0 .00 RPM |
| YRI_NA19118 | 0 .00 RPM |
| YRI_NA19213 | 0 .00 RPM |
| YRI_NA19214 | 0 .00 RPM |
| YRI_NA19223 | 0 .00 RPM |