RetrogeneDB ID: | retro_hsap_30 | ||
Retrocopylocation | Organism: | Human (Homo sapiens) | |
| Coordinates: | 7:4899527..4901438(-) | ||
| Located in intron of: | ENSG00000157927 | ||
Retrocopyinformation | Ensembl ID: | ENSG00000218823 | |
| Aliases: | PAPOLB, PAPT, TPAP | ||
| Status: | KNOWN_PROTEIN_CODING | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | PAPOLA | ||
| Ensembl ID: | ENSG00000090060 | ||
| Aliases: | PAPOLA, PAP | ||
| Description: | poly(A) polymerase alpha [Source:HGNC Symbol;Acc:14981] |

Retrocopy-Parental alignment summary:
>retro_hsap_30
ATGCCGTTTCCGGTGACAACCCAGGGACCACCGCAGCCGGCGCCGCCGCCGAATCGCTACGGCGTCTCCTCGCCTATCAG
TCTAGCGGTCCCCAAGGAGACGGACTGCCTCCTCACCCAGAGGCTAATAGAAACCCTCAGGCCCTTCGGGGTCTTCGAAG
AGGAAGAGGAACTGCAGCGCAGGATTTTAGTTTTGGAAAAATTAAATAATCTGGTAAAGGAATGGATACGCGAAATCAGT
GAAAGCAAGAGTCTTCCCCAGTCTGTAATTGAAAACGTTGGAGGAAAGATTTTTACGTTTGGCTCTTACAGATTAGGAGT
ACATACGAAAGGCGCAGATATTGACGCCTTGTGCGTTGCACCAAGTCATGTGGATCGAAGCGACTTTTTCACCTCATTCT
ATGCTAAACTGAAACTACAGGAGGAAGTGAAAGATTTAAGGGCTGTCGAGGAGGCATTTGTGCCAGTTATCAAACTGTGT
TTTGATGGGATAGAGATTGATATTTTATTTGCAAGATTAGCACTACAGACTATTCCAGAAGATTTGGACTTAAGAGATGA
CAGTCTACTTAAAAATTTAGACATTAGATGCATAAGAAGCCTTAATGGTTGCCGGGTAACCGATGAAATTTTACATCTAG
TGCCAAACATTGACAACTTCAGGCTGACTCTGAGAGCCATCAAACTGTGGGCCAAGTGCCACAATATCTATTCCAATATA
TTAGGTTTCCTCGGAGGTGTTTCCTGGGCCATGCTAGTAGCAAGAACTTGTCAGCTTTATCCAAATGCAGTAGCGTCAAC
TCTTGTACGGAAATTCTTCTTGGTATTTTCAGAATGGGAATGGCCAAACCCAGTGTTACTGAAGGAGCCTGAAGAACGGA
ATCTTAATTTGCCTGTATGGGACCCAAGAGTAAATCCCAGTGATAGGTACCATCTTATGCCTATCATCACACCAGCATAC
CCACAGCAGAACTCCACATACAACGTGTCTATTTCAACCAGGATGGTCATGATTGAGGAGTTTAAACAGGGGCTTGCTAT
CACACACGAGATTTTGCTAAGTAAGGCAGAGTGGTCCAAACTCTTTGAAGCTCCAAGCTTCTTTCAAAAGTACAAGCATT
ATATTGTACTTCTGGCAAGTGCATCAACAGAAAAACAACATTTAGAATGGGTGGGCTTGGTGGAATCAAAGATCCGAATC
CTGGTTGGGAGCTTGGAGAAGAATGAATTTATTACACTGGCACATGTGAATCCACAGTCATTTCCAGCACCCAAAGAAAA
TCCTGATATGGAAGAATTTCGTACAATGTGGGTGATTGGGTTAGGGCTAAAAAAGCCAGATAATTCTGAAATTCTCAGCA
TTGATCTCACCTATGATATCCAGTCTTTCACAGATACTGTTTATAGGCAAGCAGTGAATAGTAAGATGTTTGAGATGGGT
ATGAAAATTACTGCAATGCATTTAAGAAGAAAGGAACTTCACCAGCTGCTGCCTCATCATGTGCTTCAGGACAAGAAAGC
ACACTCAACAGAAGGTAGAAGATTGACAGATTTGAACGACAGCAGCTTTGACTTGTCTGCAGGCTGTGAAAACAGCATGT
CTGTGCCTTCATCTACTAGCACTATGAAGACAGGCCCATTGATTAGCAGTTCTCAGGGTAGAAACAGTCCTGCCTTGGCT
GTAATGACAGCATCTGTGGCTAACATACAGGCTACTGAATTTTCCTTGCAACAGGTGAATACCAATGAAAGTTCAGGGGT
TGCATTAAACGAAAGTATTCCTCACGCTGTCTCTCAGCCTGCCATTTCTCCATCACCAAAGGCCATGGTCGCCAGAGTTG
TTTCTTCAACATGTCTCATAAGCCATCCAGACCTTCAGGAAACTCAGCAACAAACATATCTAATCCTATAG
ATGCCGTTTCCGGTGACAACCCAGGGACCACCGCAGCCGGCGCCGCCGCCGAATCGCTACGGCGTCTCCTCGCCTATCAG
TCTAGCGGTCCCCAAGGAGACGGACTGCCTCCTCACCCAGAGGCTAATAGAAACCCTCAGGCCCTTCGGGGTCTTCGAAG
AGGAAGAGGAACTGCAGCGCAGGATTTTAGTTTTGGAAAAATTAAATAATCTGGTAAAGGAATGGATACGCGAAATCAGT
GAAAGCAAGAGTCTTCCCCAGTCTGTAATTGAAAACGTTGGAGGAAAGATTTTTACGTTTGGCTCTTACAGATTAGGAGT
ACATACGAAAGGCGCAGATATTGACGCCTTGTGCGTTGCACCAAGTCATGTGGATCGAAGCGACTTTTTCACCTCATTCT
ATGCTAAACTGAAACTACAGGAGGAAGTGAAAGATTTAAGGGCTGTCGAGGAGGCATTTGTGCCAGTTATCAAACTGTGT
TTTGATGGGATAGAGATTGATATTTTATTTGCAAGATTAGCACTACAGACTATTCCAGAAGATTTGGACTTAAGAGATGA
CAGTCTACTTAAAAATTTAGACATTAGATGCATAAGAAGCCTTAATGGTTGCCGGGTAACCGATGAAATTTTACATCTAG
TGCCAAACATTGACAACTTCAGGCTGACTCTGAGAGCCATCAAACTGTGGGCCAAGTGCCACAATATCTATTCCAATATA
TTAGGTTTCCTCGGAGGTGTTTCCTGGGCCATGCTAGTAGCAAGAACTTGTCAGCTTTATCCAAATGCAGTAGCGTCAAC
TCTTGTACGGAAATTCTTCTTGGTATTTTCAGAATGGGAATGGCCAAACCCAGTGTTACTGAAGGAGCCTGAAGAACGGA
ATCTTAATTTGCCTGTATGGGACCCAAGAGTAAATCCCAGTGATAGGTACCATCTTATGCCTATCATCACACCAGCATAC
CCACAGCAGAACTCCACATACAACGTGTCTATTTCAACCAGGATGGTCATGATTGAGGAGTTTAAACAGGGGCTTGCTAT
CACACACGAGATTTTGCTAAGTAAGGCAGAGTGGTCCAAACTCTTTGAAGCTCCAAGCTTCTTTCAAAAGTACAAGCATT
ATATTGTACTTCTGGCAAGTGCATCAACAGAAAAACAACATTTAGAATGGGTGGGCTTGGTGGAATCAAAGATCCGAATC
CTGGTTGGGAGCTTGGAGAAGAATGAATTTATTACACTGGCACATGTGAATCCACAGTCATTTCCAGCACCCAAAGAAAA
TCCTGATATGGAAGAATTTCGTACAATGTGGGTGATTGGGTTAGGGCTAAAAAAGCCAGATAATTCTGAAATTCTCAGCA
TTGATCTCACCTATGATATCCAGTCTTTCACAGATACTGTTTATAGGCAAGCAGTGAATAGTAAGATGTTTGAGATGGGT
ATGAAAATTACTGCAATGCATTTAAGAAGAAAGGAACTTCACCAGCTGCTGCCTCATCATGTGCTTCAGGACAAGAAAGC
ACACTCAACAGAAGGTAGAAGATTGACAGATTTGAACGACAGCAGCTTTGACTTGTCTGCAGGCTGTGAAAACAGCATGT
CTGTGCCTTCATCTACTAGCACTATGAAGACAGGCCCATTGATTAGCAGTTCTCAGGGTAGAAACAGTCCTGCCTTGGCT
GTAATGACAGCATCTGTGGCTAACATACAGGCTACTGAATTTTCCTTGCAACAGGTGAATACCAATGAAAGTTCAGGGGT
TGCATTAAACGAAAGTATTCCTCACGCTGTCTCTCAGCCTGCCATTTCTCCATCACCAAAGGCCATGGTCGCCAGAGTTG
TTTCTTCAACATGTCTCATAAGCCATCCAGACCTTCAGGAAACTCAGCAACAAACATATCTAATCCTATAG
ORF - retro_hsap_30 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 86.08 % |
| Parental protein coverage: | 83.89 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | MPFPVTTQGSQQTQPPQKHYGITSPISLAAPKETDCVLTQKLIETLKPFGVFEEEEELQRRILILGKLNN |
| MPFPVTTQG..Q..PP...YG..SPISLA.PKETDC.LTQ.LIETL.PFGVFEEEEELQRRIL.L.KLNN | |
| Retrocopy | MPFPVTTQGPPQPAPPPNRYGVSSPISLAVPKETDCLLTQRLIETLRPFGVFEEEEELQRRILVLEKLNN |
| Parental | LVKEWIREISESKNLPQSVIENVGGKIFTFGSYRLGVHTKGADIDALCVAPRHVDRSDFFTSFYDKLKLQ |
| LVKEWIREISESK.LPQSVIENVGGKIFTFGSYRLGVHTKGADIDALCVAP.HVDRSDFFTSFY.KLKLQ | |
| Retrocopy | LVKEWIREISESKSLPQSVIENVGGKIFTFGSYRLGVHTKGADIDALCVAPSHVDRSDFFTSFYAKLKLQ |
| Parental | EEVKDLRAVEEAFVPVIKLCFDGIEIDILFARLALQTIPEDLDLRDDSLLKNLDIRCIRSLNGCRVTDEI |
| EEVKDLRAVEEAFVPVIKLCFDGIEIDILFARLALQTIPEDLDLRDDSLLKNLDIRCIRSLNGCRVTDEI | |
| Retrocopy | EEVKDLRAVEEAFVPVIKLCFDGIEIDILFARLALQTIPEDLDLRDDSLLKNLDIRCIRSLNGCRVTDEI |
| Parental | LHLVPNIDNFRLTLRAIKLWAKRHNIYSNILGFLGGVSWAMLVARTCQLYPNAIASTLVHKFFLVFSKWE |
| LHLVPNIDNFRLTLRAIKLWAK.HNIYSNILGFLGGVSWAMLVARTCQLYPNA.ASTLV.KFFLVFS.WE | |
| Retrocopy | LHLVPNIDNFRLTLRAIKLWAKCHNIYSNILGFLGGVSWAMLVARTCQLYPNAVASTLVRKFFLVFSEWE |
| Parental | WPNPVLLKQPEECNLNLPVWDPRVNPSDRYHLMPIITPAYPQQNSTYNVSVSTRMVMVEEFKQGLAITDE |
| WPNPVLLK.PEE.NLNLPVWDPRVNPSDRYHLMPIITPAYPQQNSTYNVS.STRMVM.EEFKQGLAIT.E | |
| Retrocopy | WPNPVLLKEPEERNLNLPVWDPRVNPSDRYHLMPIITPAYPQQNSTYNVSISTRMVMIEEFKQGLAITHE |
| Parental | ILLSKAEWSKLFEAPNFFQKYKHYIVLLASAPTEKQRLEWVGLVESKIRILVGSLEKNEFITLAHVNPQS |
| ILLSKAEWSKLFEAP.FFQKYKHYIVLLASA.TEKQ.LEWVGLVESKIRILVGSLEKNEFITLAHVNPQS | |
| Retrocopy | ILLSKAEWSKLFEAPSFFQKYKHYIVLLASASTEKQHLEWVGLVESKIRILVGSLEKNEFITLAHVNPQS |
| Parental | FPAPKENPDKEEFRTMWVIGLVFKKTENSENLSVDLTYDIQSFTDTVYRQAINSKMFEVDMKIAAMHVKR |
| FPAPKENPD.EEFRTMWVIGL..KK..NSE.LS.DLTYDIQSFTDTVYRQA.NSKMFE..MKI.AMH..R | |
| Retrocopy | FPAPKENPDMEEFRTMWVIGLGLKKPDNSEILSIDLTYDIQSFTDTVYRQAVNSKMFEMGMKITAMHLRR |
| Parental | KQLHQLLPNHVLQKKKKHSTEGVKLTALNDSSLDLSMDSDNSMSVPSPTSATKTSPLNSSGSSQGRNSPA |
| K.LHQLLP.HVLQ.KK.HSTEG..LT.LNDSS.DLS....NSMSVPS.TS..KT.PL.S..SSQGRNSPA | |
| Retrocopy | KELHQLLPHHVLQDKKAHSTEGRRLTDLNDSSFDLSAGCENSMSVPSSTSTMKTGPLIS--SSQGRNSPA |
| Parental | PAVTAASVTNIQATEVSVPQVNSSESSGGTSSESIPQTATQPAISPPPKPTVSRVVSSTRLVNPP |
| .AV..ASV.NIQATE.S..QVN..ESSG....ESIP....QPAISP.PK..V.RVVSST.L...P | |
| Retrocopy | LAVMTASVANIQATEFSLQQVNTNESSGVALNESIPHAVSQPAISPSPKAMVARVVSSTCLISHP |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| bodymap2_adipose | 0 .00 RPM | 158 .61 RPM |
| bodymap2_adrenal | 0 .06 RPM | 187 .15 RPM |
| bodymap2_brain | 0 .12 RPM | 100 .09 RPM |
| bodymap2_breast | 0 .23 RPM | 136 .85 RPM |
| bodymap2_colon | 0 .00 RPM | 264 .22 RPM |
| bodymap2_heart | 0 .04 RPM | 147 .41 RPM |
| bodymap2_kidney | 0 .00 RPM | 216 .61 RPM |
| bodymap2_liver | 0 .00 RPM | 99 .68 RPM |
| bodymap2_lung | 0 .00 RPM | 174 .75 RPM |
| bodymap2_lymph_node | 0 .00 RPM | 311 .33 RPM |
| bodymap2_ovary | 0 .06 RPM | 299 .56 RPM |
| bodymap2_prostate | 0 .00 RPM | 240 .77 RPM |
| bodymap2_skeletal_muscle | 0 .00 RPM | 93 .94 RPM |
| bodymap2_testis | 24 .79 RPM | 285 .45 RPM |
| bodymap2_thyroid | 0 .00 RPM | 188 .99 RPM |
| bodymap2_white_blood_cells | 0 .00 RPM | 355 .96 RPM |
RNA Polymerase II actvity may be related with retro_hsap_30 in 1 libraries
| ENCODE library ID | Target | ChIP-Seq Peak coordinates |
|---|---|---|
| ENCFF002CQM | POLR2A | 7:4901284..4901764 |
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_176111 | 1816 libraries | 9 libraries | 4 libraries | 0 libraries | 0 libraries |
| TSS #2 | TSS_176112 | 1747 libraries | 75 libraries | 4 libraries | 1 library | 2 libraries |
The graphical summary, for retro_hsap_30 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1829 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
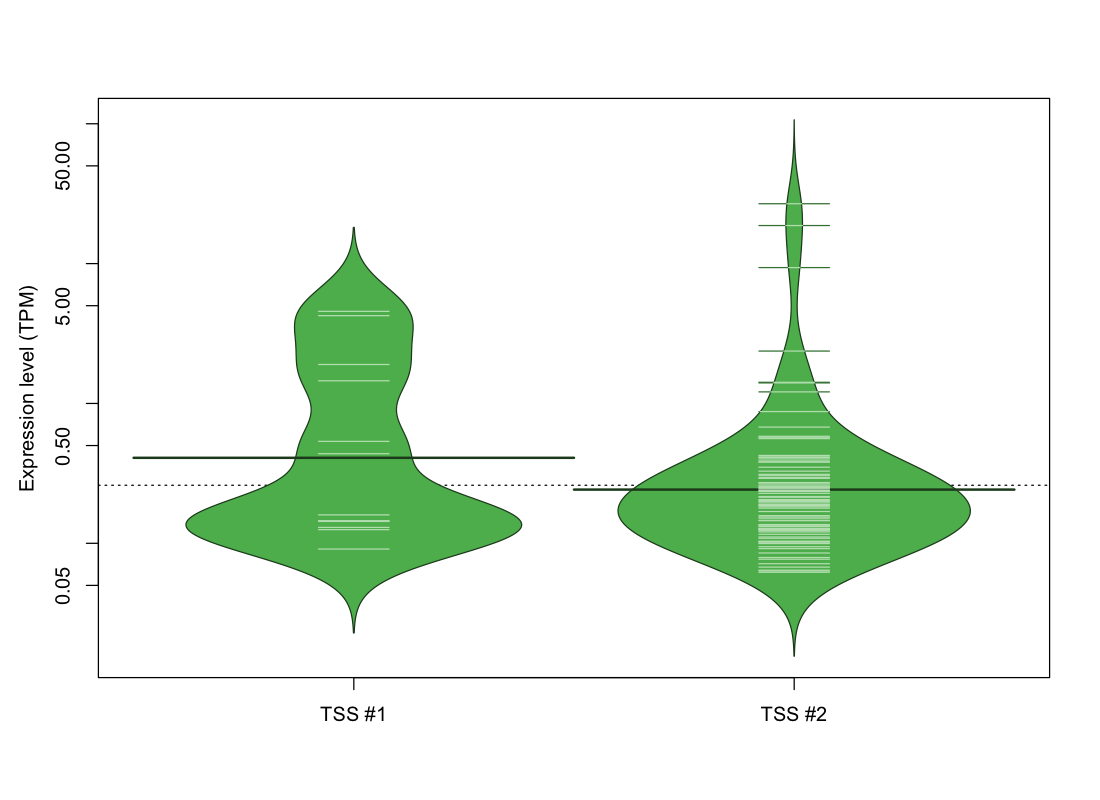
retro_hsap_30 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_hsap_30 has 9 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 20 parental genes, and 22 retrocopies.Expression level across human populations :
| Library | Retrogene expression |
|---|---|
| CEU_NA11831 | 0 .00 RPM |
| CEU_NA11843 | 0 .00 RPM |
| CEU_NA11930 | 0 .00 RPM |
| CEU_NA12004 | 0 .00 RPM |
| CEU_NA12400 | 0 .00 RPM |
| CEU_NA12751 | 0 .02 RPM |
| CEU_NA12760 | 0 .00 RPM |
| CEU_NA12827 | 0 .00 RPM |
| CEU_NA12872 | 0 .00 RPM |
| CEU_NA12873 | 0 .00 RPM |
| FIN_HG00183 | 0 .00 RPM |
| FIN_HG00277 | 0 .00 RPM |
| FIN_HG00315 | 0 .03 RPM |
| FIN_HG00321 | 0 .06 RPM |
| FIN_HG00328 | 0 .00 RPM |
| FIN_HG00338 | 0 .00 RPM |
| FIN_HG00349 | 0 .03 RPM |
| FIN_HG00375 | 0 .00 RPM |
| FIN_HG00377 | 0 .00 RPM |
| FIN_HG00378 | 0 .00 RPM |
| GBR_HG00099 | 0 .00 RPM |
| GBR_HG00111 | 0 .00 RPM |
| GBR_HG00114 | 0 .00 RPM |
| GBR_HG00119 | 0 .00 RPM |
| GBR_HG00131 | 0 .00 RPM |
| GBR_HG00133 | 0 .00 RPM |
| GBR_HG00134 | 0 .00 RPM |
| GBR_HG00137 | 0 .00 RPM |
| GBR_HG00142 | 0 .00 RPM |
| GBR_HG00143 | 0 .00 RPM |
| TSI_NA20512 | 0 .00 RPM |
| TSI_NA20513 | 0 .00 RPM |
| TSI_NA20518 | 0 .00 RPM |
| TSI_NA20532 | 0 .03 RPM |
| TSI_NA20538 | 0 .00 RPM |
| TSI_NA20756 | 0 .00 RPM |
| TSI_NA20765 | 0 .00 RPM |
| TSI_NA20771 | 0 .00 RPM |
| TSI_NA20786 | 0 .00 RPM |
| TSI_NA20798 | 0 .00 RPM |
| YRI_NA18870 | 0 .00 RPM |
| YRI_NA18907 | 0 .00 RPM |
| YRI_NA18916 | 0 .00 RPM |
| YRI_NA19093 | 0 .00 RPM |
| YRI_NA19099 | 0 .00 RPM |
| YRI_NA19114 | 0 .00 RPM |
| YRI_NA19118 | 0 .00 RPM |
| YRI_NA19213 | 0 .00 RPM |
| YRI_NA19214 | 0 .00 RPM |
| YRI_NA19223 | 0 .00 RPM |