RetrogeneDB ID: | retro_mmus_159 | ||
Retrocopy location | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 4:86575918..86576860(+) | ||
| Located in intron of: | None | ||
Retrocopy information | Ensembl ID: | ENSMUSG00000070934 | |
| Aliases: | Rraga, 1300010C19Rik, AI255374, FIP-1, RAGA | ||
| Status: | KNOWN_PROTEIN_CODING | ||
Parental gene information | Parental gene summary: | ||
| Parental gene symbol: | Rragb | ||
| Ensembl ID: | ENSMUSG00000041658 | ||
| Aliases: | None | ||
| Description: | Ras-related GTP binding B [Source:MGI Symbol;Acc:MGI:3038613] |

Retrocopy-Parental alignment summary:
>retro_mmus_159
ATGCCCAATACAGCCATGAAGAAAAAGGTGCTGTTGATGGGAAAGAGCGGGTCGGGGAAGACCAGCATGAGGTCGATCAT
CTTTGCAAATTATATTGCGCGCGACACTAGGCGCCTGGGTGCCACCATCGACGTGGAGCACTCCCACGTCCGATTCTTGG
GGAACCTGGTGCTGAACCTGTGGGACTGTGGCGGCCAGGACACCTTTATGGAAAATTACTTCACCAGCCAGCGGGACAAC
ATCTTCCGTAATGTGGAGGTTCTGATATACGTGTTTGACGTAGAGAGCCGCGAACTGGAAAAAGATATGCATTATTACCA
ATCGTGTCTGGAAGCCATCCTCCAGAATTCACCTGACGCCAAAATCTTCTGCCTGGTGCACAAAATGGATCTGGTTCAGG
AAGATCAACGTGACCTGATTTTTAAAGAGCGAGAGGAAGACCTGAGGCGTTTGTCTCGCCCGCTGGAGTGCGCTTGTTTT
CGAACGTCCATCTGGGATGAGACGCTCTACAAAGCCTGGTCCAGCATCGTCTATCAGCTGATCCCCAACGTTCAGCAGCT
GGAGATGAACCTAAGGAATTTTGCCCAGATTATTGAGGCTGATGAAGTTCTGCTGTTCGAGAGAGCTACGTTCTTGGTGA
TTTCCCACTACCAGTGCAAAGAGCAGCGAGATGTGCACCGCTTTGAGAAGATCAGCAACATCATCAAGCAGTTTAAGCTG
AGCTGCAGTAAATTGGCCGCTTCTTTCCAGAGCATGGAAGTGAGGAATTCTAACTTCGCTGCCTTCATCGACATCTTTAC
ATCAAACACGTACGTGATGGTGGTCATGTCAGATCCATCCATCCCTTCTGCAGCTACTCTGATCAACATTCGTAATGCCA
GGAAACACTTTGAGAAGCTAGAAAGAGTAGATGGGCCCAAGCACAGCCTCCTCATGCGTTGA
ORF - retro_mmus_159 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 97.43 % |
| Parental protein coverage: | 72.73 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected: | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | TFDVEHSHVRFLGNLVLNLWDCGGQDTFMENYFTSQRDNIFRNVEVLIYVFDVESRELEKDMHYYQSCLE |
| T.DVEHSHVRFLGNLVLNLWDCGGQDTFMENYFTSQRDNIFRNVEVLIYVFDVESRELEKDMHYYQSCLE | |
| Retrocopy | TIDVEHSHVRFLGNLVLNLWDCGGQDTFMENYFTSQRDNIFRNVEVLIYVFDVESRELEKDMHYYQSCLE |
| Parental | AILQNSPEAKIFCLVHKMDLVQEDQRDLIFKEREEDLRRLSRPLECSCFRTSIWDETLYKAWSSIVYQLI |
| AILQNSP.AKIFCLVHKMDLVQEDQRDLIFKEREEDLRRLSRPLEC.CFRTSIWDETLYKAWSSIVYQLI | |
| Retrocopy | AILQNSPDAKIFCLVHKMDLVQEDQRDLIFKEREEDLRRLSRPLECACFRTSIWDETLYKAWSSIVYQLI |
| Parental | PNVQQLEMNLRNFAEIIEADEVLLFERATFLVISHYQCKEQRDAHRFEKISNIIKQFKLSCSKLAASFQS |
| PNVQQLEMNLRNFA.IIEADEVLLFERATFLVISHYQCKEQRD.HRFEKISNIIKQFKLSCSKLAASFQS | |
| Retrocopy | PNVQQLEMNLRNFAQIIEADEVLLFERATFLVISHYQCKEQRDVHRFEKISNIIKQFKLSCSKLAASFQS |
| Parental | MEVRNSNFAAFIDIFTSNTYVMVVMSDPSIPSAATLINIRNARKHFEKLERVDGPKQCLLMR |
| MEVRNSNFAAFIDIFTSNTYVMVVMSDPSIPSAATLINIRNARKHFEKLERVDGPK..LLMR | |
| Retrocopy | MEVRNSNFAAFIDIFTSNTYVMVVMSDPSIPSAATLINIRNARKHFEKLERVDGPKHSLLMR |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 66 .63 RPM | 55 .04 RPM |
| SRP007412_cerebellum | 65 .83 RPM | 55 .71 RPM |
| SRP007412_heart | 28 .36 RPM | 2 .77 RPM |
| SRP007412_kidney | 30 .66 RPM | 2 .62 RPM |
| SRP007412_liver | 19 .91 RPM | 0 .14 RPM |
| SRP007412_testis | 13 .13 RPM | 7 .50 RPM |
RNA Polymerase II activity may be related with retro_mmus_159 in 3 libraries
| ENCODE library ID | Target | ChIP-Seq Peak coordinates |
|---|---|---|
| ENCFF001XVA | POLR2A | 4:86575320..86576152 |
| ENCFF001YIJ | POLR2A | 4:86574905..86578992 |
| ENCFF001YKA | POLR2A | 4:86574943..86579071 |
6 EST(s) were mapped to retro_mmus_159 retrocopy
| EST ID | Start | End | Identity | Match | Mis-match | Score |
|---|---|---|---|---|---|---|
| AA673540 | 86576223 | 86576550 | 99.1 | 324 | 3 | 321 |
| AA690642 | 86576035 | 86576235 | 99.5 | 199 | 0 | 197 |
| AW259959 | 86575920 | 86576167 | 96 | 237 | 10 | 227 |
| BF123878 | 86576063 | 86576391 | 97.3 | 326 | 2 | 321 |
| CV676679 | 86576136 | 86576394 | 100 | 257 | 0 | 256 |
| CX212840 | 86576036 | 86576356 | 98.5 | 320 | 0 | 318 |
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_106688 | 19 libraries | 1 library | 87 libraries | 282 libraries | 683 libraries |
| TSS #2 | TSS_106689 | 4 libraries | 0 libraries | 0 libraries | 3 libraries | 1065 libraries |
| TSS #3 | TSS_106690 | 40 libraries | 10 libraries | 629 libraries | 366 libraries | 27 libraries |
| TSS #4 | TSS_106691 | 71 libraries | 137 libraries | 740 libraries | 117 libraries | 7 libraries |
| TSS #5 | TSS_106692 | 298 libraries | 529 libraries | 232 libraries | 12 libraries | 1 library |
| TSS #6 | TSS_106693 | 492 libraries | 447 libraries | 109 libraries | 15 libraries | 9 libraries |
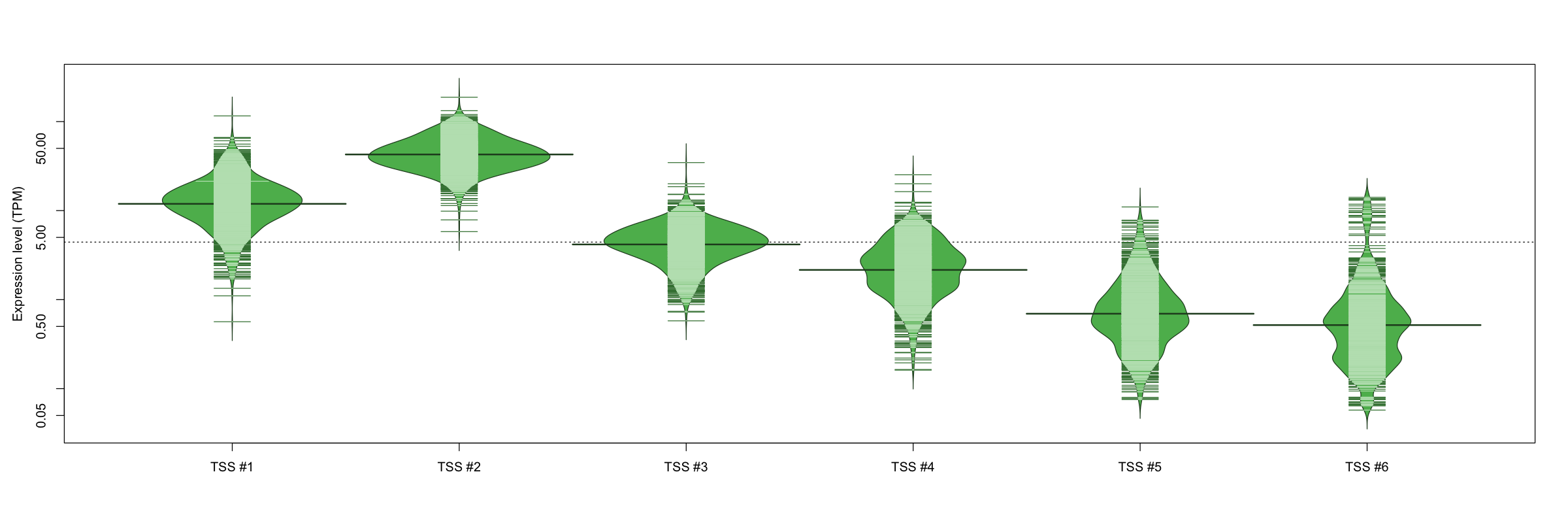
The graphical summary, for retro_mmus_159 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
retro_mmus_159 was not experimentally validated.