RetrogeneDB ID: | retro_hsap_56 | ||
Retrocopylocation | Organism: | Human (Homo sapiens) | |
| Coordinates: | 1:241797097..241799068(-) | ||
| Located in intron of: | ENSG00000054277 | ||
Retrocopyinformation | Ensembl ID: | ENSG00000203668 | |
| Aliases: | CHML, REP2 | ||
| Status: | KNOWN_PROTEIN_CODING | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | CHM | ||
| Ensembl ID: | ENSG00000188419 | ||
| Aliases: | CHM, DXS540, GGTA, HSD-32, REP-1, TCD | ||
| Description: | choroideremia (Rab escort protein 1) [Source:HGNC Symbol;Acc:1940] |

Retrocopy-Parental alignment summary:
>retro_hsap_56
ATGGCGGACAATCTTCCCACAGAGTTTGATGTGGTTATAATAGGGACAGGTTTGCCCGAATCCATCCTTGCAGCTGCATG
TTCAAGAAGTGGTCAGAGGGTTCTGCATATTGATTCAAGAAGCTACTATGGAGGAAACTGGGCTAGTTTCAGCTTTTCAG
GATTGCTATCCTGGTTGAAGGAGTATCAGCAAAACAATGACATTGGGGAAGAAAGTACTGTTGTATGGCAGGACCTGATC
CATGAAACAGAAGAAGCCATCACTCTTCGCAAGAAGGATGAAACTATTCAACACACAGAAGCTTTTTGCTACGCCAGTCA
GGATATGGAGGACAACGTTGAAGAGATTGGTGCTCTGCAGAAAAATCCTTCTTTGGGGGTGTCTAATACCTTCACTGAAG
TTCTGGATTCTGCATTACCTGAAGAAAGCCAGTTATCGTATTTTAATAGCGACGAAATGCCTGCAAAACACACTCAGAAA
AGTGATACAGAGATTTCACTAGAAGTAACTGATGTAGAGGAATCAGTGGAGAAGGAAAAGTATTGTGGAGATAAAACTTG
TATGCACACAGTTTCAGATAAAGATGGAGATAAAGATGAAAGCAAATCTACAGTAGAAGATAAGGCCGATGAACCAATTA
GAAATAGGATTACTTACTCTCAAATAGTTAAAGAAGGCAGGAGGTTTAATATTGATTTGGTGTCAAAACTGCTGTATTCT
CAAGGATTGCTAATTGATCTTTTAATCAAATCAGATGTTAGTCGTTATGTAGAATTTAAAAATGTCACTAGGATTCTTGC
ATTTCGGGAAGGAAAGGTAGAACAAGTTCCTTGTTCCAGAGCAGATGTCTTTAATAGCAAGGAACTCACCATGGTTGAAA
AGAGGATGCTAATGAAATTTCTCACATTTTGTTTAGAGTATGAACAACATCCTGATGAATACCAAGCTTTCAGGCAGTGT
TCATTTTCAGAATACTTAAAAACTAAAAAACTAACTCCCAACCTTCAACATTTTGTACTGCACTCAATTGCAATGACATC
AGAATCATCTTGCACTACAATAGATGGTCTTAACGCAACTAAAAACTTCCTTCAGTGTCTCGGACGGTTTGGCAACACCC
CCTTTTTATTTCCCTTGTATGGCCAAGGAGAAATTCCCCAGGGTTTCTGTAGGATGTGTGCAGTTTTTGGTGGAATCTAT
TGTCTTCGTCATAAAGTACAATGCTTTGTAGTCGACAAAGAATCTGGACGATGTAAAGCAATTATAGATCACTTTGGTCA
AAGAATAAATGCTAAATATTTTATTGTGGAAGACAGTTACCTTTCTGAGGAAACATGCTCAAATGTGCAGTATAAGCAGA
TCTCTAGGGCAGTACTCATTACAGATCAGTCTATACTAAAGACAGATTTAGATCAGCAGACTTCCATTCTGATAGTTCCT
CCAGCAGAGCCAGGAGCTTGTGCTGTACGGGTCACAGAATTATGTTCTTCAACCATGACATGCATGAAGGACACCTATCT
GGTACATTTGACATGTTCATCTTCTAAAACAGCAAGAGAAGACTTAGAATCAGTGGTGAAGAAATTATTCACTCCGTATA
CTGAAACAGAAATAAACGAGGAAGAACTTACAAAGCCAAGACTCTTGTGGGCTCTTTATTTTAATATGAGAGATTCCTCG
GGAATCAGCAGAAGCTCGTATAATGGCTTGCCTTCCAATGTTTATGTCTGCTCTGGGCCTGACTGTGGCCTGGGAAATGA
GCATGCTGTCAAGCAAGCTGAAACACTTTTCCAGGAGATCTTTCCAACTGAAGAATTCTGCCCTCCACCTCCAAATCCAG
AAGACATTATCTTTGATGGTGATGATAAGCAGCCAGAGGCTCCTGGAACCAATAATGTAGTAATGGCCAAACTAGAATCC
TCTGAGGAAAGCAAAAACCTAGAAAGCCCAGAGAAGCACCTTCAAAATTAG
ATGGCGGACAATCTTCCCACAGAGTTTGATGTGGTTATAATAGGGACAGGTTTGCCCGAATCCATCCTTGCAGCTGCATG
TTCAAGAAGTGGTCAGAGGGTTCTGCATATTGATTCAAGAAGCTACTATGGAGGAAACTGGGCTAGTTTCAGCTTTTCAG
GATTGCTATCCTGGTTGAAGGAGTATCAGCAAAACAATGACATTGGGGAAGAAAGTACTGTTGTATGGCAGGACCTGATC
CATGAAACAGAAGAAGCCATCACTCTTCGCAAGAAGGATGAAACTATTCAACACACAGAAGCTTTTTGCTACGCCAGTCA
GGATATGGAGGACAACGTTGAAGAGATTGGTGCTCTGCAGAAAAATCCTTCTTTGGGGGTGTCTAATACCTTCACTGAAG
TTCTGGATTCTGCATTACCTGAAGAAAGCCAGTTATCGTATTTTAATAGCGACGAAATGCCTGCAAAACACACTCAGAAA
AGTGATACAGAGATTTCACTAGAAGTAACTGATGTAGAGGAATCAGTGGAGAAGGAAAAGTATTGTGGAGATAAAACTTG
TATGCACACAGTTTCAGATAAAGATGGAGATAAAGATGAAAGCAAATCTACAGTAGAAGATAAGGCCGATGAACCAATTA
GAAATAGGATTACTTACTCTCAAATAGTTAAAGAAGGCAGGAGGTTTAATATTGATTTGGTGTCAAAACTGCTGTATTCT
CAAGGATTGCTAATTGATCTTTTAATCAAATCAGATGTTAGTCGTTATGTAGAATTTAAAAATGTCACTAGGATTCTTGC
ATTTCGGGAAGGAAAGGTAGAACAAGTTCCTTGTTCCAGAGCAGATGTCTTTAATAGCAAGGAACTCACCATGGTTGAAA
AGAGGATGCTAATGAAATTTCTCACATTTTGTTTAGAGTATGAACAACATCCTGATGAATACCAAGCTTTCAGGCAGTGT
TCATTTTCAGAATACTTAAAAACTAAAAAACTAACTCCCAACCTTCAACATTTTGTACTGCACTCAATTGCAATGACATC
AGAATCATCTTGCACTACAATAGATGGTCTTAACGCAACTAAAAACTTCCTTCAGTGTCTCGGACGGTTTGGCAACACCC
CCTTTTTATTTCCCTTGTATGGCCAAGGAGAAATTCCCCAGGGTTTCTGTAGGATGTGTGCAGTTTTTGGTGGAATCTAT
TGTCTTCGTCATAAAGTACAATGCTTTGTAGTCGACAAAGAATCTGGACGATGTAAAGCAATTATAGATCACTTTGGTCA
AAGAATAAATGCTAAATATTTTATTGTGGAAGACAGTTACCTTTCTGAGGAAACATGCTCAAATGTGCAGTATAAGCAGA
TCTCTAGGGCAGTACTCATTACAGATCAGTCTATACTAAAGACAGATTTAGATCAGCAGACTTCCATTCTGATAGTTCCT
CCAGCAGAGCCAGGAGCTTGTGCTGTACGGGTCACAGAATTATGTTCTTCAACCATGACATGCATGAAGGACACCTATCT
GGTACATTTGACATGTTCATCTTCTAAAACAGCAAGAGAAGACTTAGAATCAGTGGTGAAGAAATTATTCACTCCGTATA
CTGAAACAGAAATAAACGAGGAAGAACTTACAAAGCCAAGACTCTTGTGGGCTCTTTATTTTAATATGAGAGATTCCTCG
GGAATCAGCAGAAGCTCGTATAATGGCTTGCCTTCCAATGTTTATGTCTGCTCTGGGCCTGACTGTGGCCTGGGAAATGA
GCATGCTGTCAAGCAAGCTGAAACACTTTTCCAGGAGATCTTTCCAACTGAAGAATTCTGCCCTCCACCTCCAAATCCAG
AAGACATTATCTTTGATGGTGATGATAAGCAGCCAGAGGCTCCTGGAACCAATAATGTAGTAATGGCCAAACTAGAATCC
TCTGAGGAAAGCAAAAACCTAGAAAGCCCAGAGAAGCACCTTCAAAATTAG
ORF - retro_hsap_56 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 71.98 % |
| Parental protein coverage: | 99.54 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | MADTLPSEFDVIVIGTGLPESIIAAACSRSGRRVLHVDSRSYYGGNWASFSFSGLLSWLKEYQENSDIVS |
| MAD.LP.EFDV..IGTGLPESI.AAACSRSG.RVLH.DSRSYYGGNWASFSFSGLLSWLKEYQ.N.DI.. | |
| Retrocopy | MADNLPTEFDVVIIGTGLPESILAAACSRSGQRVLHIDSRSYYGGNWASFSFSGLLSWLKEYQQNNDIGE |
| Parental | DSPV-WQDQILENEEAIALSRKDKTIQHVEVFCYASQDLHEDVEEAGALQKNHALVTSANSTEAADSAFL |
| .S.V.WQD.I.E.EEAI.L..KD.TIQH.E.FCYASQD....VEE.GALQKN..L..S...TE..DSA.L | |
| Retrocopy | ESTVVWQDLIHETEEAITLRKKDETIQHTEAFCYASQDMEDNVEEIGALQKNPSLGVSNTFTEVLDSA-L |
| Parental | PTEDESLSTMSCEMLTEQTPSSDPENALEVNGAEVTGEKENHCDDKTCVPSTSAED--MSENVPIAEDTT |
| P.E.......S.EM....T..SD.E..LEV...E...EKE..C.DKTC....S..D....E.....ED.. | |
| Retrocopy | PEESQLSYFNSDEMPAKHTQKSDTEISLEVTDVEESVEKEKYCGDKTCMHTVSDKDGDKDESKSTVEDKA |
| Parental | EQPKKNRITYSQIIKEGRRFNIDLVSKLLYSRGLLIDLLIKSNVSRYAEFKNITRILAFREGRVEQVPCS |
| ..P..NRITYSQI.KEGRRFNIDLVSKLLYS.GLLIDLLIKS.VSRY.EFKN.TRILAFREG.VEQVPCS | |
| Retrocopy | DEPIRNRITYSQIVKEGRRFNIDLVSKLLYSQGLLIDLLIKSDVSRYVEFKNVTRILAFREGKVEQVPCS |
| Parental | RADVFNSKQLTMVEKRMLMKFLTFCMEYEKYPDEYKGYEEITFYEYLKTQKLTPNLQYIVMHSIAMTSET |
| RADVFNSK.LTMVEKRMLMKFLTFC.EYE..PDEY.......F.EYLKT.KLTPNLQ..V.HSIAMTSE. | |
| Retrocopy | RADVFNSKELTMVEKRMLMKFLTFCLEYEQHPDEYQAFRQCSFSEYLKTKKLTPNLQHFVLHSIAMTSES |
| Parental | ASSTIDGLKATKNFLHCLGRYGNTPFLFPLYGQGELPQCFCRMCAVFGGIYCLRHSVQCLVVDKESRKCK |
| ...TIDGL.ATKNFL.CLGR.GNTPFLFPLYGQGE.PQ.FCRMCAVFGGIYCLRH.VQC.VVDKES..CK | |
| Retrocopy | SCTTIDGLNATKNFLQCLGRFGNTPFLFPLYGQGEIPQGFCRMCAVFGGIYCLRHKVQCFVVDKESGRCK |
| Parental | AIIDQFGQRIISEHFLVEDSYFPENMCSRVQYRQISRAVLITDRSVLKTDSDQQISILTVPAEEPGTFAV |
| AIID.FGQRI....F.VEDSY..E..CS.VQY.QISRAVLITD.S.LKTD.DQQ.SIL.VP..EPG..AV | |
| Retrocopy | AIIDHFGQRINAKYFIVEDSYLSEETCSNVQYKQISRAVLITDQSILKTDLDQQTSILIVPPAEPGACAV |
| Parental | RVIELCSSTMTCMKGTYLVHLTCTSSKTAREDLESVVQKLFVPYTEMEIENEQVEKPRILWALYFNMRDS |
| RV.ELCSSTMTCMK.TYLVHLTC.SSKTAREDLESVV.KLF.PYTE.EI..E...KPR.LWALYFNMRDS | |
| Retrocopy | RVTELCSSTMTCMKDTYLVHLTCSSSKTAREDLESVVKKLFTPYTETEINEEELTKPRLLWALYFNMRDS |
| Parental | SDISRSCYNDLPSNVYVCSGPDCGLGNDNAVKQAETLFQEICPNEDFCPPPPNPEDIILDGDSLQPEASE |
| S.ISRS.YN.LPSNVYVCSGPDCGLGN..AVKQAETLFQEI.P.E.FCPPPPNPEDII.DGD..QPEA.. | |
| Retrocopy | SGISRSSYNGLPSNVYVCSGPDCGLGNEHAVKQAETLFQEIFPTEEFCPPPPNPEDIIFDGDDKQPEAPG |
| Parental | SSAIPEANSETFKESTNLGNLEE |
| ......A..E...ES.NL...E. | |
| Retrocopy | TNNVVMAKLESSEESKNLESPEK |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| bodymap2_adipose | 4 .81 RPM | 19 .92 RPM |
| bodymap2_adrenal | 1 .47 RPM | 20 .69 RPM |
| bodymap2_brain | 6 .00 RPM | 43 .88 RPM |
| bodymap2_breast | 2 .55 RPM | 25 .10 RPM |
| bodymap2_colon | 4 .10 RPM | 19 .28 RPM |
| bodymap2_heart | 3 .03 RPM | 19 .73 RPM |
| bodymap2_kidney | 3 .39 RPM | 40 .93 RPM |
| bodymap2_liver | 0 .21 RPM | 11 .58 RPM |
| bodymap2_lung | 1 .74 RPM | 20 .89 RPM |
| bodymap2_lymph_node | 3 .44 RPM | 20 .92 RPM |
| bodymap2_ovary | 9 .08 RPM | 24 .46 RPM |
| bodymap2_prostate | 2 .32 RPM | 33 .15 RPM |
| bodymap2_skeletal_muscle | 2 .16 RPM | 17 .71 RPM |
| bodymap2_testis | 10 .43 RPM | 21 .21 RPM |
| bodymap2_thyroid | 5 .33 RPM | 40 .69 RPM |
| bodymap2_white_blood_cells | 8 .35 RPM | 38 .53 RPM |
RNA Polymerase II actvity near the 5' end of retro_hsap_56 was not detected
14 EST(s) were mapped to retro_hsap_56 retrocopy
| EST ID | Start | End | Identity | Match | Mis-match | Score |
|---|---|---|---|---|---|---|
| AA669486 | 241797016 | 241797447 | 98.4 | 422 | 7 | 413 |
| AL708330 | 241797752 | 241798058 | 98.1 | 304 | 0 | 300 |
| AU123990 | 241797031 | 241797820 | 99.5 | 783 | 2 | 777 |
| AU137744 | 241797713 | 241798416 | 97.9 | 692 | 3 | 680 |
| AU139019 | 241797829 | 241798439 | 97.7 | 605 | 0 | 596 |
| BP267405 | 241797700 | 241798298 | 97.4 | 581 | 16 | 564 |
| BP268285 | 241797736 | 241798298 | 99.9 | 561 | 1 | 560 |
| BP282990 | 241797241 | 241797823 | 100 | 581 | 0 | 581 |
| BU569726 | 241797236 | 241797697 | 99 | 456 | 5 | 451 |
| BX474169 | 241797056 | 241797544 | 100 | 488 | 0 | 488 |
| BX474181 | 241798254 | 241798538 | 99 | 277 | 3 | 274 |
| CF139504 | 241797545 | 241798085 | 99.9 | 539 | 1 | 538 |
| N31339 | 241797571 | 241797708 | 100 | 132 | 0 | 132 |
| R91881 | 241797090 | 241797487 | 97.2 | 388 | 1 | 380 |
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_91775 | 1112 libraries | 621 libraries | 94 libraries | 2 libraries | 0 libraries |
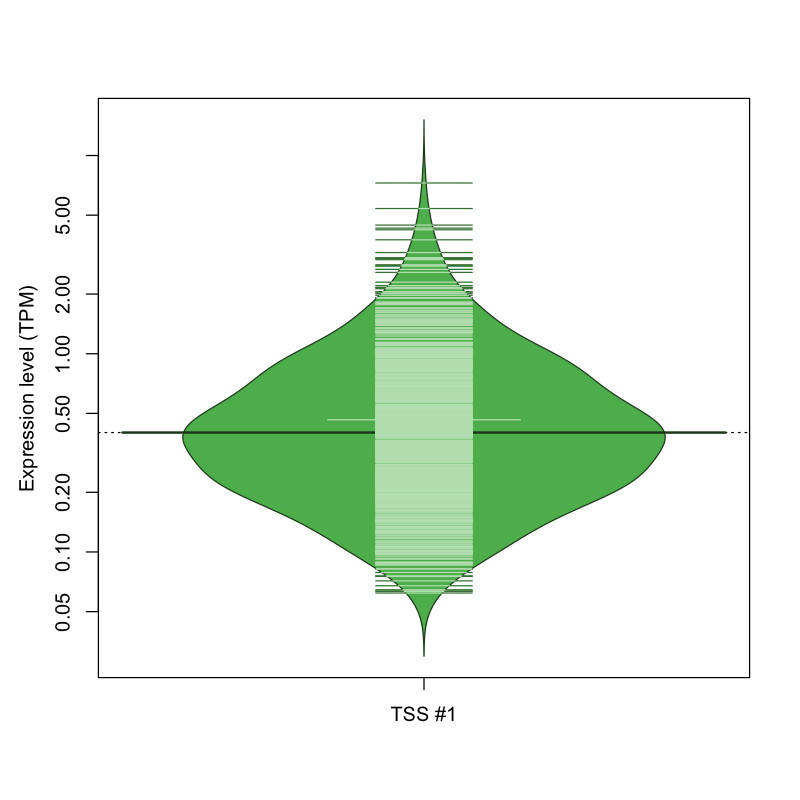
The graphical summary, for retro_hsap_56 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1829 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
retro_hsap_56 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_hsap_56 has 10 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 21 parental genes, and 21 retrocopies.Expression level across human populations :
| Library | Retrogene expression |
|---|---|
| CEU_NA11831 | 4 .64 RPM |
| CEU_NA11843 | 2 .33 RPM |
| CEU_NA11930 | 11 .11 RPM |
| CEU_NA12004 | 5 .58 RPM |
| CEU_NA12400 | 2 .95 RPM |
| CEU_NA12751 | 9 .18 RPM |
| CEU_NA12760 | 3 .73 RPM |
| CEU_NA12827 | 5 .66 RPM |
| CEU_NA12872 | 7 .59 RPM |
| CEU_NA12873 | 6 .57 RPM |
| FIN_HG00183 | 8 .59 RPM |
| FIN_HG00277 | 7 .17 RPM |
| FIN_HG00315 | 6 .21 RPM |
| FIN_HG00321 | 9 .40 RPM |
| FIN_HG00328 | 4 .47 RPM |
| FIN_HG00338 | 5 .43 RPM |
| FIN_HG00349 | 4 .72 RPM |
| FIN_HG00375 | 6 .31 RPM |
| FIN_HG00377 | 4 .29 RPM |
| FIN_HG00378 | 7 .27 RPM |
| GBR_HG00099 | 12 .87 RPM |
| GBR_HG00111 | 3 .20 RPM |
| GBR_HG00114 | 6 .86 RPM |
| GBR_HG00119 | 11 .54 RPM |
| GBR_HG00131 | 12 .16 RPM |
| GBR_HG00133 | 5 .35 RPM |
| GBR_HG00134 | 9 .83 RPM |
| GBR_HG00137 | 4 .01 RPM |
| GBR_HG00142 | 7 .14 RPM |
| GBR_HG00143 | 7 .69 RPM |
| TSI_NA20512 | 3 .59 RPM |
| TSI_NA20513 | 20 .97 RPM |
| TSI_NA20518 | 7 .33 RPM |
| TSI_NA20532 | 8 .98 RPM |
| TSI_NA20538 | 8 .94 RPM |
| TSI_NA20756 | 4 .77 RPM |
| TSI_NA20765 | 9 .78 RPM |
| TSI_NA20771 | 6 .67 RPM |
| TSI_NA20786 | 9 .18 RPM |
| TSI_NA20798 | 8 .26 RPM |
| YRI_NA18870 | 5 .21 RPM |
| YRI_NA18907 | 5 .65 RPM |
| YRI_NA18916 | 6 .57 RPM |
| YRI_NA19093 | 3 .93 RPM |
| YRI_NA19099 | 6 .52 RPM |
| YRI_NA19114 | 7 .47 RPM |
| YRI_NA19118 | 5 .86 RPM |
| YRI_NA19213 | 8 .08 RPM |
| YRI_NA19214 | 10 .77 RPM |
| YRI_NA19223 | 11 .74 RPM |