RetrogeneDB ID: | retro_hsap_3976 | ||
Retrocopylocation | Organism: | Human (Homo sapiens) | |
| Coordinates: | 8:81213203..81213713(+) | ||
| Located in intron of: | ENSG00000253238 | ||
Retrocopyinformation | Ensembl ID: | ENSG00000213791 | |
| Aliases: | None | ||
| Status: | KNOWN_PSEUDOGENE | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | RPS5 | ||
| Ensembl ID: | ENSG00000083845 | ||
| Aliases: | RPS5, S5 | ||
| Description: | ribosomal protein S5 [Source:HGNC Symbol;Acc:10426] |

Retrocopy-Parental alignment summary:
>retro_hsap_3976
TTGCAGGATTACACTGCAGTGAAGGAGAAGTATGCCAAGTACCTGCCTCACAGTGCAGAGCGGTATGCCGCCAAACGCTT
CCGCAAAGCTCAGTGCCCCATCGTGGAGTGCCTCACTTACTCCATGATGACGCACGGCCGCAACAACCGCAAGAAGCTCA
TGATCATGCGCATCGTCAAGCACGCCTTCGAGATCATCCAGCTGCTCACAGGCGACAACCCTCTGCAGGTCCTGGTGAAC
GCCATCATCAACAGTGGTCCCCGGGAGGACTCCACACTCACTGGGCGAGCCAGGATCGTGAGACGACAGGCTGTGGCCGT
GTCCCCATTAAGCCGTGTGAATCAGGCCATCTGGCCGCAGTGCACAGGAGCTCTTGAGGCTGCCTTCCGGAACATCAAGA
CCATTGCTGAGTTCCTGGCAGATGAGCTCATCAGTGCTGTCAAGGGCTCCTCCAACTCCTATGCTAGCAAGAAGAAGGAC
GAGCTTGAGTGTGTGGCCAAGTCCAACCGC
TTGCAGGATTACACTGCAGTGAAGGAGAAGTATGCCAAGTACCTGCCTCACAGTGCAGAGCGGTATGCCGCCAAACGCTT
CCGCAAAGCTCAGTGCCCCATCGTGGAGTGCCTCACTTACTCCATGATGACGCACGGCCGCAACAACCGCAAGAAGCTCA
TGATCATGCGCATCGTCAAGCACGCCTTCGAGATCATCCAGCTGCTCACAGGCGACAACCCTCTGCAGGTCCTGGTGAAC
GCCATCATCAACAGTGGTCCCCGGGAGGACTCCACACTCACTGGGCGAGCCAGGATCGTGAGACGACAGGCTGTGGCCGT
GTCCCCATTAAGCCGTGTGAATCAGGCCATCTGGCCGCAGTGCACAGGAGCTCTTGAGGCTGCCTTCCGGAACATCAAGA
CCATTGCTGAGTTCCTGGCAGATGAGCTCATCAGTGCTGTCAAGGGCTCCTCCAACTCCTATGCTAGCAAGAAGAAGGAC
GAGCTTGAGTGTGTGGCCAAGTCCAACCGC
ORF - retro_hsap_3976 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 85.88 % |
| Parental protein coverage: | 83.33 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | LQDYIAVKEKYAKYLPHSAGRYAAKRFRKAQCPIVERLTNSMMMHGRNNGKKLMTVRIVKHAFEIIHLLT |
| LQDY.AVKEKYAKYLPHSA.RYAAKRFRKAQCPIVE.LT.SMM.HGRNN.KKLM..RIVKHAFEII.LLT | |
| Retrocopy | LQDYTAVKEKYAKYLPHSAERYAAKRFRKAQCPIVECLTYSMMTHGRNNRKKLMIMRIVKHAFEIIQLLT |
| Parental | GENPLQVLVNAIINSGPREDSTRIGRAGTVRRQAVDVSPLRRVNQAIWLLCTGAREAAFRNIKTIAECLA |
| G.NPLQVLVNAIINSGPREDST..GRA..VRRQAV.VSPL.RVNQAIW..CTGA.EAAFRNIKTIAE.LA | |
| Retrocopy | GDNPLQVLVNAIINSGPREDSTLTGRARIVRRQAVAVSPLSRVNQAIWPQCTGALEAAFRNIKTIAEFLA |
| Parental | DELINAAKGSSNSYAIKKKDELERVAKSNR |
| DELI.A.KGSSNSYA.KKKDELE.VAKSNR | |
| Retrocopy | DELISAVKGSSNSYASKKKDELECVAKSNR |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| bodymap2_adipose | 0 .00 RPM | 249 .17 RPM |
| bodymap2_adrenal | 0 .02 RPM | 279 .29 RPM |
| bodymap2_brain | 0 .02 RPM | 59 .63 RPM |
| bodymap2_breast | 1 .52 RPM | 186 .71 RPM |
| bodymap2_colon | 0 .00 RPM | 374 .15 RPM |
| bodymap2_heart | 0 .00 RPM | 83 .35 RPM |
| bodymap2_kidney | 0 .00 RPM | 138 .43 RPM |
| bodymap2_liver | 0 .00 RPM | 138 .78 RPM |
| bodymap2_lung | 0 .00 RPM | 323 .07 RPM |
| bodymap2_lymph_node | 0 .00 RPM | 384 .86 RPM |
| bodymap2_ovary | 0 .00 RPM | 416 .21 RPM |
| bodymap2_prostate | 0 .00 RPM | 457 .87 RPM |
| bodymap2_skeletal_muscle | 0 .00 RPM | 239 .88 RPM |
| bodymap2_testis | 0 .00 RPM | 195 .62 RPM |
| bodymap2_thyroid | 0 .00 RPM | 197 .43 RPM |
| bodymap2_white_blood_cells | 0 .00 RPM | 395 .35 RPM |
RNA Polymerase II actvity near the 5' end of retro_hsap_3976 was not detected
No EST(s) were mapped for retro_hsap_3976 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_186651 | 1279 libraries | 455 libraries | 77 libraries | 11 libraries | 7 libraries |
The graphical summary, for retro_hsap_3976 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1829 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
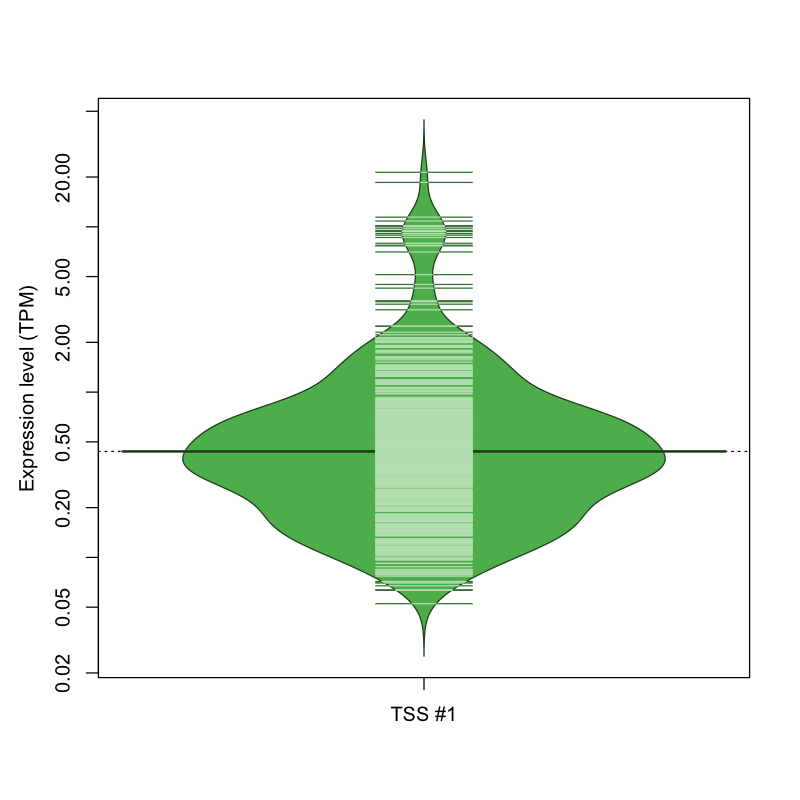
retro_hsap_3976 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_hsap_3976 has 2 orthologous retrocopies within eutheria group .| Species | RetrogeneDB ID |
|---|---|
| Pongo abelii | retro_pabe_3270 |
| Macaca mulatta | retro_mmul_77 |
Parental genes homology:
Parental genes homology involve 18 parental genes, and 49 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Canis familiaris | ENSCAFG00000002366 | 1 retrocopy | |
| Callithrix jacchus | ENSCJAG00000000304 | 6 retrocopies | |
| Echinops telfairi | ENSETEG00000013543 | 1 retrocopy | |
| Homo sapiens | ENSG00000083845 | 5 retrocopies | |
| Gorilla gorilla | ENSGGOG00000004258 | 4 retrocopies | |
| Macropus eugenii | ENSMEUG00000015400 | 3 retrocopies | |
| Myotis lucifugus | ENSMLUG00000012641 | 1 retrocopy | |
| Macaca mulatta | ENSMMUG00000008729 | 3 retrocopies | |
| Mustela putorius furo | ENSMPUG00000008308 | 1 retrocopy | |
| Ornithorhynchus anatinus | ENSOANG00000011921 | 1 retrocopy | |
| Otolemur garnettii | ENSOGAG00000013088 | 2 retrocopies | |
| Procavia capensis | ENSPCAG00000011034 | 2 retrocopies | |
| Pongo abelii | ENSPPYG00000010500 | 7 retrocopies | |
| Pan troglodytes | ENSPTRG00000011591 | 6 retrocopies | |
| Ictidomys tridecemlineatus | ENSSTOG00000005935 | 1 retrocopy | |
| Tupaia belangeri | ENSTBEG00000016955 | 2 retrocopies | |
| Tarsius syrichta | ENSTSYG00000000076 | 2 retrocopies | |
| Tursiops truncatus | ENSTTRG00000001991 | 1 retrocopy |
Expression level across human populations :
| Library | Retrogene expression |
|---|---|
| CEU_NA11831 | 0 .02 RPM |
| CEU_NA11843 | 0 .00 RPM |
| CEU_NA11930 | 0 .03 RPM |
| CEU_NA12004 | 0 .04 RPM |
| CEU_NA12400 | 0 .00 RPM |
| CEU_NA12751 | 0 .00 RPM |
| CEU_NA12760 | 0 .04 RPM |
| CEU_NA12827 | 0 .00 RPM |
| CEU_NA12872 | 0 .00 RPM |
| CEU_NA12873 | 0 .00 RPM |
| FIN_HG00183 | 0 .00 RPM |
| FIN_HG00277 | 0 .00 RPM |
| FIN_HG00315 | 0 .00 RPM |
| FIN_HG00321 | 0 .00 RPM |
| FIN_HG00328 | 0 .00 RPM |
| FIN_HG00338 | 0 .00 RPM |
| FIN_HG00349 | 0 .00 RPM |
| FIN_HG00375 | 0 .00 RPM |
| FIN_HG00377 | 0 .00 RPM |
| FIN_HG00378 | 0 .00 RPM |
| GBR_HG00099 | 0 .00 RPM |
| GBR_HG00111 | 0 .00 RPM |
| GBR_HG00114 | 0 .00 RPM |
| GBR_HG00119 | 0 .00 RPM |
| GBR_HG00131 | 0 .00 RPM |
| GBR_HG00133 | 0 .00 RPM |
| GBR_HG00134 | 0 .00 RPM |
| GBR_HG00137 | 0 .03 RPM |
| GBR_HG00142 | 0 .00 RPM |
| GBR_HG00143 | 0 .00 RPM |
| TSI_NA20512 | 0 .00 RPM |
| TSI_NA20513 | 0 .00 RPM |
| TSI_NA20518 | 0 .06 RPM |
| TSI_NA20532 | 0 .00 RPM |
| TSI_NA20538 | 0 .00 RPM |
| TSI_NA20756 | 0 .00 RPM |
| TSI_NA20765 | 0 .02 RPM |
| TSI_NA20771 | 0 .00 RPM |
| TSI_NA20786 | 0 .03 RPM |
| TSI_NA20798 | 0 .00 RPM |
| YRI_NA18870 | 0 .00 RPM |
| YRI_NA18907 | 0 .00 RPM |
| YRI_NA18916 | 0 .00 RPM |
| YRI_NA19093 | 0 .00 RPM |
| YRI_NA19099 | 0 .00 RPM |
| YRI_NA19114 | 0 .05 RPM |
| YRI_NA19118 | 0 .02 RPM |
| YRI_NA19213 | 0 .05 RPM |
| YRI_NA19214 | 0 .00 RPM |
| YRI_NA19223 | 0 .04 RPM |