RetrogeneDB ID: | retro_hsap_15 | ||
Retrocopylocation | Organism: | Human (Homo sapiens) | |
| Coordinates: | 6:28227149..28228358(+) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | ENSG00000189134 | |
| Aliases: | NKAPL, C6orf194, bA424I5.1 | ||
| Status: | KNOWN_PROTEIN_CODING | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | NKAP | ||
| Ensembl ID: | ENSG00000101882 | ||
| Aliases: | None | ||
| Description: | NFKB activating protein [Source:HGNC Symbol;Acc:29873] |

Retrocopy-Parental alignment summary:
>retro_hsap_15
ATGCCCCCAGTATCCCGGTCCAGCTATTCCGAGGACATCGTGGGCTCTCGGAGAAGGCGACGCAGCTCCTCGGGGAGCCC
ACCATCCCCGCAGAGCAGATGTTCCTCTTGGGATGGCTGTTCCCGCTCTCACTCCCGCGGCCGTGAGGGCCTCAGGCCTC
CTTGGAGTGAGTTGGACGTGGGCGCTCTTTACCCCTTTAGTCGCTCTGGGTCGCGAGGGCGGCTCCCAAGATTCCGCAAC
TACGCCTTCGCGTCCTCCTGGTCGACCTCGTATAGTGGATATCGCTACCATCGTCACTGCTATGCAGAAGAACGGCAGTC
AGCGGAAGACTACGAGAAGGAAGAGAGCCATCGGCAGAGGAGGCTGAAGGAGAGAGAGAGGATTGGGGAATTGGGAGCGC
CTGAAGTGTGGGGGCCGTCTCCAAAGTTCCCTCAGCTAGATTCTGACGAACATACCCCAGTTGAGGATGAAGAAGAGGTA
ACGCATCAGAAAAGCAGCAGTTCAGATTCCAACTCGGAAGAACATAGGAAAAAGAAGACCAGTCGTTCAAGAAACAAGAA
AAAAAGAAAGAATAAGTCGTCTAAAAGAAAGCATAGGAAATATTCTGATAGTGACAGTAACTCAGAGTCTGACACAAATT
CTGACTCTGATGATGATAAAAAGAGAGTTAAAGCCAAGAAGAAAAAGAAGAAAAAGAAACACAAAACAAAGAAAAAGAAG
AATAAGAAAACCAAAAAAGAATCCAGTGACTCAAGCTGTAAAGACTCAGAAGAGGACTTGTCAGAAGCTACCTGGATGGA
GCAGCCAAATGTGGCAGATACTATGGATTTAATAGGGCCAGAAGCACCTATAATACATACCTCTCAAGATGAAAAACCTT
TGAAGTATGGCCATGCTTTGCTTCCCGGTGAAGGTGCAGCTATGGCTGAGTATGTAAAAGCTGGAAAGCGAATCCCACGA
AGAGGTGAAATTGGGTTGACAAGTGAAGAGATCGGTTCTTTTGAATGCTCAGGTTATGTCATGAGTGGTAGCAGGCATCG
CAGAATGGAGGCTGTACGACTGCGTAAGGAGAACCAGATCTACAGTGCTGATGAGAAGAGAGCTCTTGCATCCTTTAACC
AAGAAGAGAGACGAAAGAGAGAAAGTAAGATTTTAGCCAGTTTCCGAGAGATGGTGCACAAAAAGACAAAAGAGAAAGAT
GACAAGTAA
ATGCCCCCAGTATCCCGGTCCAGCTATTCCGAGGACATCGTGGGCTCTCGGAGAAGGCGACGCAGCTCCTCGGGGAGCCC
ACCATCCCCGCAGAGCAGATGTTCCTCTTGGGATGGCTGTTCCCGCTCTCACTCCCGCGGCCGTGAGGGCCTCAGGCCTC
CTTGGAGTGAGTTGGACGTGGGCGCTCTTTACCCCTTTAGTCGCTCTGGGTCGCGAGGGCGGCTCCCAAGATTCCGCAAC
TACGCCTTCGCGTCCTCCTGGTCGACCTCGTATAGTGGATATCGCTACCATCGTCACTGCTATGCAGAAGAACGGCAGTC
AGCGGAAGACTACGAGAAGGAAGAGAGCCATCGGCAGAGGAGGCTGAAGGAGAGAGAGAGGATTGGGGAATTGGGAGCGC
CTGAAGTGTGGGGGCCGTCTCCAAAGTTCCCTCAGCTAGATTCTGACGAACATACCCCAGTTGAGGATGAAGAAGAGGTA
ACGCATCAGAAAAGCAGCAGTTCAGATTCCAACTCGGAAGAACATAGGAAAAAGAAGACCAGTCGTTCAAGAAACAAGAA
AAAAAGAAAGAATAAGTCGTCTAAAAGAAAGCATAGGAAATATTCTGATAGTGACAGTAACTCAGAGTCTGACACAAATT
CTGACTCTGATGATGATAAAAAGAGAGTTAAAGCCAAGAAGAAAAAGAAGAAAAAGAAACACAAAACAAAGAAAAAGAAG
AATAAGAAAACCAAAAAAGAATCCAGTGACTCAAGCTGTAAAGACTCAGAAGAGGACTTGTCAGAAGCTACCTGGATGGA
GCAGCCAAATGTGGCAGATACTATGGATTTAATAGGGCCAGAAGCACCTATAATACATACCTCTCAAGATGAAAAACCTT
TGAAGTATGGCCATGCTTTGCTTCCCGGTGAAGGTGCAGCTATGGCTGAGTATGTAAAAGCTGGAAAGCGAATCCCACGA
AGAGGTGAAATTGGGTTGACAAGTGAAGAGATCGGTTCTTTTGAATGCTCAGGTTATGTCATGAGTGGTAGCAGGCATCG
CAGAATGGAGGCTGTACGACTGCGTAAGGAGAACCAGATCTACAGTGCTGATGAGAAGAGAGCTCTTGCATCCTTTAACC
AAGAAGAGAGACGAAAGAGAGAAAGTAAGATTTTAGCCAGTTTCCGAGAGATGGTGCACAAAAAGACAAAAGAGAAAGAT
GACAAGTAA
ORF - retro_hsap_15 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 69.1 % |
| Parental protein coverage: | 81.93 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | SRSRSRERPSAPRGIPFASASSSVYYGSYSRPYGSDKPWPSLLDKEREESLRQKRLSERERIGELGAPEV |
| SRS.SR.R....R...FAS..S..Y.G.............S..D.E.EES.RQ.RL.ERERIGELGAPEV | |
| Retrocopy | SRSGSRGRLPRFRNYAFASSWSTSYSGYRYHRHCYAEERQSAEDYEKEESHRQRRLKERERIGELGAPEV |
| Parental | WGLSPKNPEPDSDEHTPVEDEEP---KKSTTSASTSEEEKKKKSSRSKERSKKRRKKKSSKRKHKKYSED |
| WG.SPK.P..DSDEHTPVEDEE.....KS..S.S.SEE..KKK.SRS..R.KK.RK.KSSKRKH.KYS.D | |
| Retrocopy | WGPSPKFPQLDSDEHTPVEDEEEVTHQKSSSSDSNSEEHRKKKTSRS--RNKKKRKNKSSKRKHRKYS-D |
| Parental | SDSDSDSETDSSDEDNKRRAKKAKKKEKKKKHRSKKYKKKRSKKSRKESSDSSSKESQEEFLENPWKDRT |
| SDS.S.S.T.S...D.K.R.K.AKKK.KKKKH.....KKK..KK..KESSDSS.K.S.E...E..W.... | |
| Retrocopy | SDSNSESDTNSDSDDDKKRVK-AKKKKKKKKHKT---KKKKNKKTKKESSDSSCKDSEEDLSEATWMEQP |
| Parental | KAEEPSDLIGPEAPKTLTSQDDKPLNYGHALLPGEGAAMAEYVKAGKRIPRRGEIGLTSEEIASFECSGY |
| ......DLIGPEAP...TSQD.KPL.YGHALLPGEGAAMAEYVKAGKRIPRRGEIGLTSEEI.SFECSGY | |
| Retrocopy | NVADTMDLIGPEAPIIHTSQDEKPLKYGHALLPGEGAAMAEYVKAGKRIPRRGEIGLTSEEIGSFECSGY |
| Parental | VMSGSRHRRMEAVRLRKENQIYSADEKRALASFNQEERRKRENKILASFREMVYRKTKGKDDK |
| VMSGSRHRRMEAVRLRKENQIYSADEKRALASFNQEERRKRE.KILASFREMV..KTK.KDDK | |
| Retrocopy | VMSGSRHRRMEAVRLRKENQIYSADEKRALASFNQEERRKRESKILASFREMVHKKTKEKDDK |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| bodymap2_adipose | 2 .33 RPM | 12 .90 RPM |
| bodymap2_adrenal | 1 .64 RPM | 16 .38 RPM |
| bodymap2_brain | 2 .11 RPM | 17 .72 RPM |
| bodymap2_breast | 4 .57 RPM | 11 .87 RPM |
| bodymap2_colon | 2 .55 RPM | 16 .32 RPM |
| bodymap2_heart | 1 .88 RPM | 6 .66 RPM |
| bodymap2_kidney | 1 .63 RPM | 13 .75 RPM |
| bodymap2_liver | 0 .13 RPM | 10 .22 RPM |
| bodymap2_lung | 0 .18 RPM | 11 .27 RPM |
| bodymap2_lymph_node | 0 .49 RPM | 14 .24 RPM |
| bodymap2_ovary | 1 .60 RPM | 20 .72 RPM |
| bodymap2_prostate | 2 .30 RPM | 16 .44 RPM |
| bodymap2_skeletal_muscle | 5 .37 RPM | 9 .40 RPM |
| bodymap2_testis | 43 .27 RPM | 19 .19 RPM |
| bodymap2_thyroid | 4 .69 RPM | 14 .20 RPM |
| bodymap2_white_blood_cells | 0 .28 RPM | 17 .36 RPM |
RNA Polymerase II actvity may be related with retro_hsap_15 in 4 libraries
| ENCODE library ID | Target | ChIP-Seq Peak coordinates |
|---|---|---|
| ENCFF002CHO | POLR2A | 6:28226873..28227222 |
| ENCFF002CQI | POLR2A | 6:28226824..28227260 |
| ENCFF002CQM | POLR2A | 6:28226747..28227227 |
| ENCFF002CQO | POLR2A | 6:28226821..28227351 |
34 EST(s) were mapped to retro_hsap_15 retrocopy
| EST ID | Start | End | Identity | Match | Mis-match | Score |
|---|---|---|---|---|---|---|
| AA193476 | 28227267 | 28227825 | 99.1 | 540 | 2 | 531 |
| AA426031 | 28227497 | 28227826 | 100 | 329 | 0 | 329 |
| AA719187 | 28227448 | 28227827 | 100 | 378 | 0 | 377 |
| AA889640 | 28227347 | 28227703 | 99.8 | 354 | 1 | 352 |
| AA993479 | 28227394 | 28227706 | 99.7 | 311 | 1 | 310 |
| AI633740 | 28227369 | 28227827 | 99.8 | 457 | 1 | 456 |
| AI825371 | 28227451 | 28227827 | 100 | 376 | 0 | 376 |
| AI825424 | 28227566 | 28227856 | 99 | 290 | 0 | 289 |
| AI970107 | 28227290 | 28227827 | 98.7 | 533 | 4 | 528 |
| AI989981 | 28227300 | 28227827 | 99.7 | 525 | 2 | 523 |
| AI129164 | 28227396 | 28227835 | 100 | 438 | 0 | 437 |
| AI183972 | 28227359 | 28227833 | 100 | 474 | 0 | 474 |
| AJ574107 | 28227325 | 28227696 | 99.8 | 370 | 1 | 369 |
| AW003978 | 28227238 | 28227827 | 99.7 | 587 | 2 | 585 |
| AW196436 | 28227357 | 28227827 | 98.8 | 469 | 1 | 466 |
| AW207887 | 28227306 | 28227835 | 99.7 | 524 | 2 | 519 |
| AW515746 | 28227326 | 28227827 | 99.9 | 500 | 1 | 499 |
| BE042106 | 28227473 | 28227714 | 99.2 | 230 | 2 | 226 |
| BF061938 | 28227363 | 28227827 | 99 | 459 | 5 | 454 |
| BF222720 | 28227311 | 28227827 | 99.9 | 515 | 1 | 514 |
| BF592882 | 28227224 | 28227827 | 97.2 | 588 | 13 | 571 |
| BQ186305 | 28227168 | 28227714 | 99.5 | 542 | 0 | 541 |
| BQ186490 | 28227177 | 28227805 | 99.6 | 626 | 0 | 625 |
| BX115857 | 28227115 | 28227764 | 99.9 | 647 | 1 | 646 |
| CK299257 | 28227168 | 28227714 | 99.9 | 542 | 1 | 541 |
| DB454384 | 28227091 | 28227560 | 99.8 | 468 | 1 | 467 |
| DB471905 | 28227074 | 28227570 | 99.8 | 495 | 1 | 494 |
| DB473910 | 28227074 | 28227561 | 99.6 | 485 | 2 | 483 |
| DB478241 | 28227074 | 28227566 | 99.6 | 490 | 2 | 488 |
| EL737170 | 28227149 | 28227795 | 99.7 | 641 | 2 | 639 |
| H86864 | 28227745 | 28228025 | 97.5 | 279 | 0 | 276 |
| HY019101 | 28227157 | 28227608 | 100 | 451 | 0 | 451 |
| HY202334 | 28227467 | 28227872 | 98.8 | 401 | 2 | 398 |
| W78197 | 28227097 | 28227526 | 98.6 | 420 | 2 | 410 |
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_163202 | 897 libraries | 253 libraries | 539 libraries | 111 libraries | 29 libraries |
| TSS #2 | TSS_163203 | 1620 libraries | 196 libraries | 11 libraries | 2 libraries | 0 libraries |
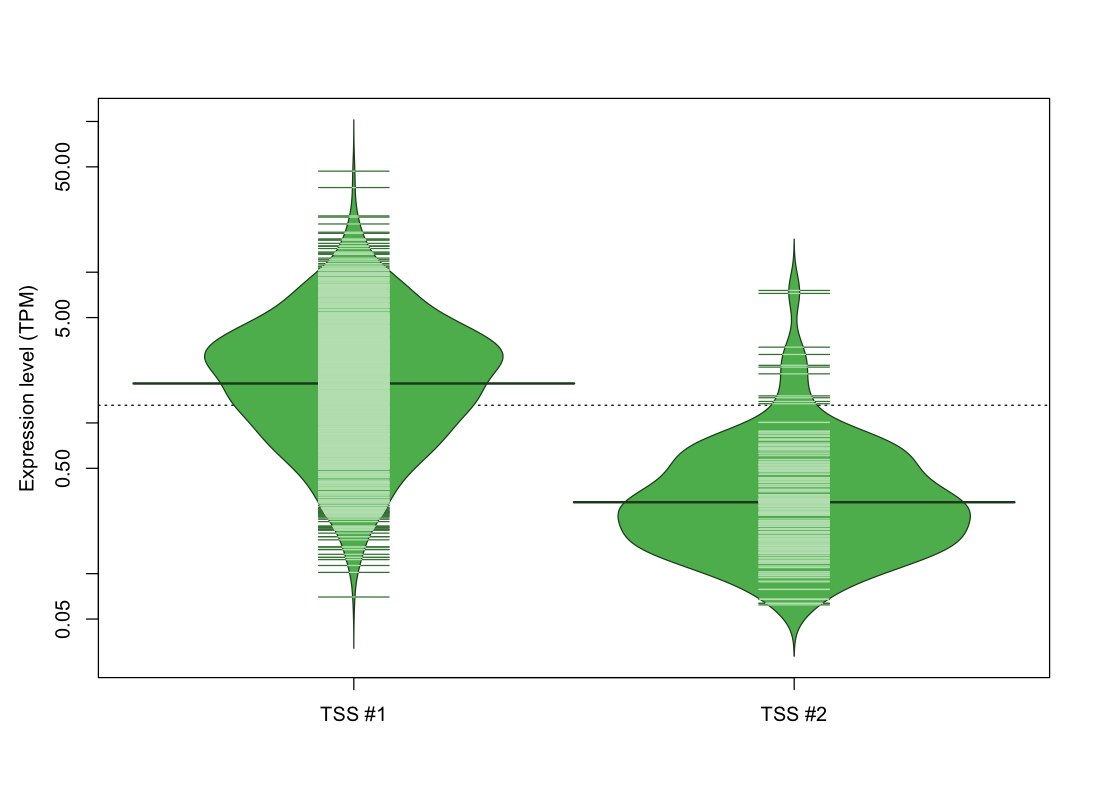
The graphical summary, for retro_hsap_15 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1829 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
retro_hsap_15 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_hsap_15 has 5 orthologous retrocopies within eutheria group .| Species | RetrogeneDB ID |
|---|---|
| Pan troglodytes | retro_ptro_117 |
| Pongo abelii | retro_pabe_198 |
| Macaca mulatta | retro_mmul_121 |
| Mus musculus | retro_mmus_279 |
| Rattus norvegicus | retro_rnor_93 |
Parental genes homology:
Parental genes homology involve 15 parental genes, and 15 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Canis familiaris | ENSCAFG00000018433 | 1 retrocopy | |
| Dasypus novemcinctus | ENSDNOG00000000072 | 1 retrocopy | |
| Echinops telfairi | ENSETEG00000001007 | 1 retrocopy | |
| Felis catus | ENSFCAG00000004392 | 1 retrocopy | |
| Homo sapiens | ENSG00000101882 | 1 retrocopy |
retro_hsap_15 ,
|
| Loxodonta africana | ENSLAFG00000002962 | 1 retrocopy | |
| Macaca mulatta | ENSMMUG00000012382 | 1 retrocopy | |
| Mus musculus | ENSMUSG00000016409 | 1 retrocopy | |
| Nomascus leucogenys | ENSNLEG00000014922 | 1 retrocopy | |
| Otolemur garnettii | ENSOGAG00000001004 | 1 retrocopy | |
| Pongo abelii | ENSPPYG00000020679 | 1 retrocopy | |
| Pan troglodytes | ENSPTRG00000022233 | 1 retrocopy | |
| Rattus norvegicus | ENSRNOG00000031169 | 1 retrocopy | |
| Sus scrofa | ENSSSCG00000012613 | 1 retrocopy | |
| Ictidomys tridecemlineatus | ENSSTOG00000001134 | 1 retrocopy |
Expression level across human populations :
| Library | Retrogene expression |
|---|---|
| CEU_NA11831 | 0 .00 RPM |
| CEU_NA11843 | 0 .06 RPM |
| CEU_NA11930 | 0 .00 RPM |
| CEU_NA12004 | 0 .00 RPM |
| CEU_NA12400 | 0 .11 RPM |
| CEU_NA12751 | 0 .05 RPM |
| CEU_NA12760 | 0 .09 RPM |
| CEU_NA12827 | 0 .05 RPM |
| CEU_NA12872 | 0 .00 RPM |
| CEU_NA12873 | 0 .03 RPM |
| FIN_HG00183 | 0 .27 RPM |
| FIN_HG00277 | 0 .11 RPM |
| FIN_HG00315 | 0 .44 RPM |
| FIN_HG00321 | 0 .09 RPM |
| FIN_HG00328 | 0 .05 RPM |
| FIN_HG00338 | 0 .13 RPM |
| FIN_HG00349 | 0 .28 RPM |
| FIN_HG00375 | 0 .24 RPM |
| FIN_HG00377 | 0 .25 RPM |
| FIN_HG00378 | 0 .13 RPM |
| GBR_HG00099 | 0 .03 RPM |
| GBR_HG00111 | 0 .21 RPM |
| GBR_HG00114 | 0 .21 RPM |
| GBR_HG00119 | 0 .43 RPM |
| GBR_HG00131 | 0 .46 RPM |
| GBR_HG00133 | 0 .31 RPM |
| GBR_HG00134 | 0 .24 RPM |
| GBR_HG00137 | 0 .11 RPM |
| GBR_HG00142 | 0 .42 RPM |
| GBR_HG00143 | 0 .16 RPM |
| TSI_NA20512 | 0 .14 RPM |
| TSI_NA20513 | 0 .02 RPM |
| TSI_NA20518 | 0 .19 RPM |
| TSI_NA20532 | 0 .10 RPM |
| TSI_NA20538 | 0 .14 RPM |
| TSI_NA20756 | 0 .20 RPM |
| TSI_NA20765 | 1 .12 RPM |
| TSI_NA20771 | 0 .20 RPM |
| TSI_NA20786 | 0 .00 RPM |
| TSI_NA20798 | 0 .19 RPM |
| YRI_NA18870 | 0 .07 RPM |
| YRI_NA18907 | 0 .42 RPM |
| YRI_NA18916 | 0 .21 RPM |
| YRI_NA19093 | 0 .05 RPM |
| YRI_NA19099 | 0 .03 RPM |
| YRI_NA19114 | 0 .10 RPM |
| YRI_NA19118 | 0 .02 RPM |
| YRI_NA19213 | 0 .69 RPM |
| YRI_NA19214 | 0 .02 RPM |
| YRI_NA19223 | 0 .06 RPM |