RetrogeneDB ID: | retro_mmus_63 | ||
Retrocopy location | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 3:141210569..141211745(-) | ||
| Located in intron of: | None | ||
Retrocopy information | Ensembl ID: | ENSMUSG00000047674 | |
| Aliases: | Pdha2, Pdhal | ||
| Status: | KNOWN_PROTEIN_CODING | ||
Parental gene information | Parental gene summary: | ||
| Parental gene symbol: | Pdha1 | ||
| Ensembl ID: | ENSMUSG00000031299 | ||
| Aliases: | Pdha1, Pdha-1 | ||
| Description: | pyruvate dehydrogenase E1 alpha 1 [Source:MGI Symbol;Acc:MGI:97532] |

Retrocopy-Parental alignment summary:
>retro_mmus_63
ATGAGGAAAATGCTGACCGCTGTGCTGTCTCACGTATTTTCGGGAATGGTCCAAAAGCCAGCTCTCAGAGGACTGCTGTC
ATCTCTGAAGTTCTCCAACGACGCCACCTGTGACATTAAGAAATGTGACCTGTACCGGCTGGAGGAGGGCCCACCGACCT
CCACCGTGCTCACCCGAGCCGAGGCCCTCAAGTACTACCGGACCATGCAGGTAATTCGGCGCATGGAGTTGAAGGCCGAC
CAGCTGTATAAGCAGAAATTCATCCGTGGTTTCTGTCACCTGTGTGATGGGCAGGAAGCCTGCTGCGTGGGGCTGGAGGC
AGGGATAAATCCCACGGATCACGTCATCACGTCCTACCGGGCTCATGGCTTCTGCTACACGCGAGGACTGTCCGTGAAGT
CCATTCTCGCCGAGCTGACTGGACGCAAAGGAGGCTGTGCTAAAGGCAAGGGAGGCTCCATGCACATGTACGGCAAGAAC
TTCTACGGTGGCAATGGCATTGTTGGGGCCCAGGTACCCCTGGGAGCTGGTGTGGCTTTTGCCTGTAAATACCTGAAGAA
TGGTCAGGTCTGCTTGGCTTTGTACGGCGATGGTGCGGCTAACCAAGGGCAGGTATTCGAAGCATACAATATGTCAGCCT
TGTGGAAATTACCCTGTGTTTTCATCTGTGAGAATAACCTCTATGGAATGGGAACCTCCAACGAGAGATCAGCAGCCAGT
ACTGATTACCACAAGAAAGGTTTTATTATCCCCGGACTGAGGGTGAATGGGATGGATATTCTCTGTGTTCGGGAGGCAAC
CAAGTTTGCAGCTGATCACTGCAGATCTGGAAAGGGGCCCATTGTGATGGAGCTGCAGACCTACCGTTATCATGGACACA
GTATGAGCGACCCAGGGATCAGTTATCGTTCACGAGAAGAAGTTCATAACGTGAGAAGTAAGAGTGATCCTATAATGCTG
CTCCGAGAGAGAATTATCAGCAACAACCTCAGCAATATTGAAGAATTGAAAGAAATTGATGCAGATGTGAAGAAAGAGGT
GGAGGACGCAGCTCAGTTTGCTACGACTGATCCAGAACCAGCTGTGGAAGATATAGCCAATTACCTCTACCACCAAGATC
CACCTTTTGAAGTCCGTGGTGCACATAAGTGGCTCAAGTATAAGTCCCACAGTTAG
ORF - retro_mmus_63 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 76.73 % |
| Parental protein coverage: | 100.0 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected: | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | MRKMLAAV-SRVLAGSAQKPASRVLVASRNFANDATFEIKKCDLHRLEEGPPVTTVLTREDGLKYYRMMQ |
| MRKML.AV.S.V..G..QKPA.R.L..S..F.NDAT..IKKCDL.RLEEGPP..TVLTR...LKYYR.MQ | |
| Retrocopy | MRKMLTAVLSHVFSGMVQKPALRGLLSSLKFSNDATCDIKKCDLYRLEEGPPTSTVLTRAEALKYYRTMQ |
| Parental | TVRRMELKADQLYKQKIIRGFCHLCDGQEACCVGLEAGINPTDHLITAYRAHGFTFTRGLPVRAILAELT |
| ..RRMELKADQLYKQK.IRGFCHLCDGQEACCVGLEAGINPTDH.IT.YRAHGF..TRGL.V..ILAELT | |
| Retrocopy | VIRRMELKADQLYKQKFIRGFCHLCDGQEACCVGLEAGINPTDHVITSYRAHGFCYTRGLSVKSILAELT |
| Parental | GRRGGCAKGKGGSMHMYAKNFYGGNGIVGAQVPLGAGIALACKYNGKDEVCLTLYGDGAANQGQIFEAYN |
| GR.GGCAKGKGGSMHMY.KNFYGGNGIVGAQVPLGAG.A.ACKY.....VCL.LYGDGAANQGQ.FEAYN | |
| Retrocopy | GRKGGCAKGKGGSMHMYGKNFYGGNGIVGAQVPLGAGVAFACKYLKNGQVCLALYGDGAANQGQVFEAYN |
| Parental | MAALWKLPCIFICENNRYGMGTSVERAAASTDYYKRGDFIPGLRVDGMDILCVREATKFAAAYCRSGKGP |
| M.ALWKLPC.FICENN.YGMGTS.ER.AASTDY.K.G..IPGLRV.GMDILCVREATKFAA..CRSGKGP | |
| Retrocopy | MSALWKLPCVFICENNLYGMGTSNERSAASTDYHKKGFIIPGLRVNGMDILCVREATKFAADHCRSGKGP |
| Parental | ILMELQTYRYHGHSMSDPGVSYRTREEIQEVRSKSDPIMLLKDRMVNSNLASVEELKEIDVEVRKEIEDA |
| I.MELQTYRYHGHSMSDPG.SYR.REE...VRSKSDPIMLL..R....NL...EELKEID..V.KE.EDA | |
| Retrocopy | IVMELQTYRYHGHSMSDPGISYRSREEVHNVRSKSDPIMLLRERIISNNLSNIEELKEIDADVKKEVEDA |
| Parental | AQFATADPEPPLEELGYHIYSSDPPFEVRGANQWIKFKSVS |
| AQFAT.DPEP..E......Y..DPPFEVRGA..W.K.KS.S | |
| Retrocopy | AQFATTDPEPAVEDIANYLYHQDPPFEVRGAHKWLKYKSHS |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .02 RPM | 136 .25 RPM |
| SRP007412_cerebellum | 0 .00 RPM | 111 .85 RPM |
| SRP007412_heart | 0 .00 RPM | 816 .64 RPM |
| SRP007412_kidney | 0 .00 RPM | 506 .84 RPM |
| SRP007412_liver | 0 .03 RPM | 102 .02 RPM |
| SRP007412_testis | 160 .13 RPM | 4 .25 RPM |
RNA Polymerase II activity near the 5' end of retro_mmus_63 was not detected
No EST(s) were mapped for retro_mmus_63 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_92797 | 1046 libraries | 19 libraries | 2 libraries | 1 library | 4 libraries |
| TSS #2 | TSS_92798 | 1045 libraries | 21 libraries | 2 libraries | 0 libraries | 4 libraries |
| TSS #3 | TSS_92799 | 1066 libraries | 2 libraries | 3 libraries | 1 library | 0 libraries |
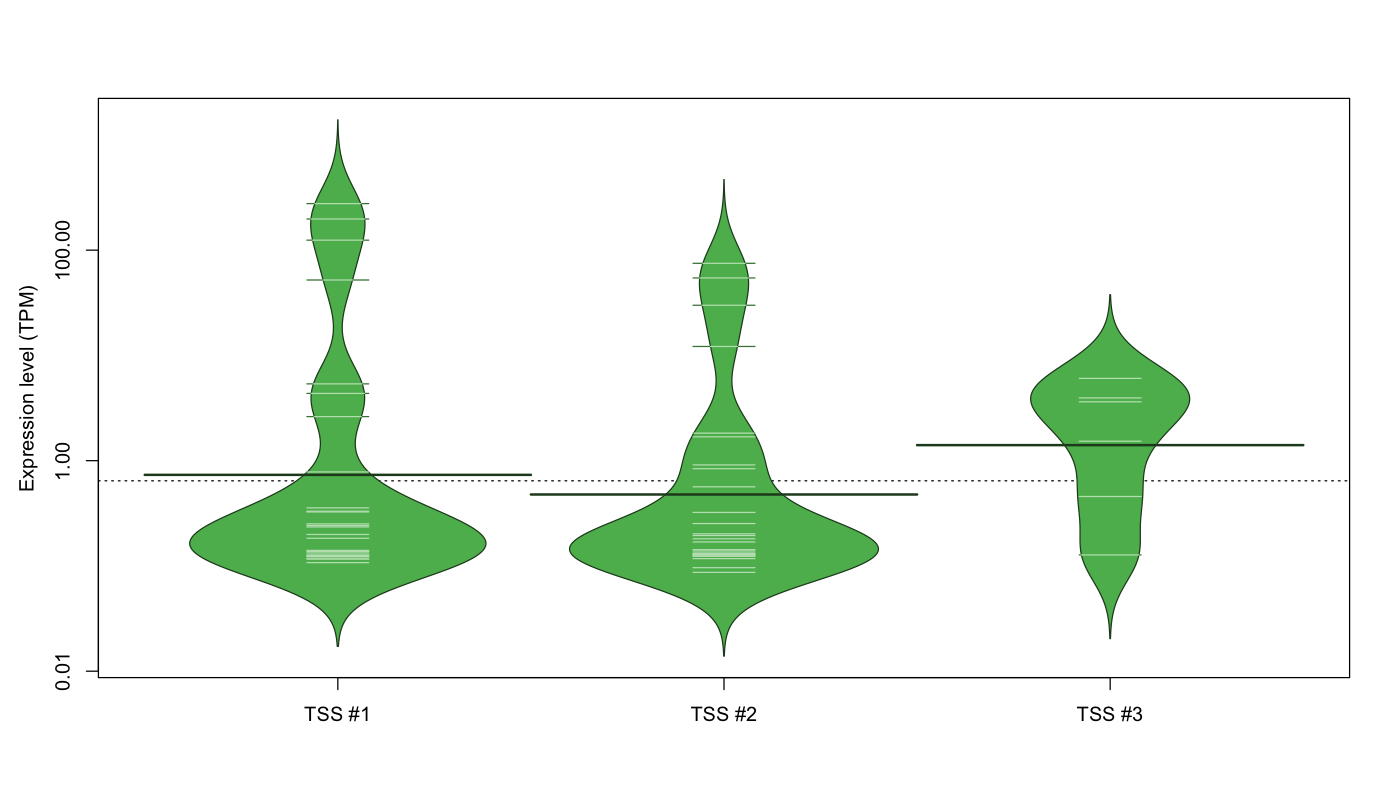
The graphical summary, for retro_mmus_63 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
retro_mmus_63 was not experimentally validated.