RetrogeneDB ID: | retro_mmus_2968 | ||
Retrocopylocation | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 7:119740127..119741096(+) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | ENSMUSG00000044341 | |
| Aliases: | None | ||
| Status: | KNOWN_PSEUDOGENE | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | Ppp1cc | ||
| Ensembl ID: | ENSMUSG00000004455 | ||
| Aliases: | Ppp1cc, PP-1G, PP1, dis2m1 | ||
| Description: | protein phosphatase 1, catalytic subunit, gamma isoform [Source:MGI Symbol;Acc:MGI:104872] |

Retrocopy-Parental alignment summary:
>retro_mmus_2968
ATGGCGGATATCGACAAACTCAACATCGACAGCATCATCCAACGGCTGCTGGAAGTGAGAGGGTCCAAGCCAGGCAAGAA
TGTCCAGCTCCAGGAGAACGAGATCCGAGGACTCTGCCTGAAGTCTCGGGAGATCTTCCTCAGTCAGCCTATCCTTTTAG
AACTTGAAGCACCACTCAAGATATGTGGTGACATCCACGGGCAGTACTATGATTTGCTCCGTCTGTTTGAATACGGTGGC
TTTCCTCCAGAGAGCAACTATTTGTTTCTCGGGGACTATGTGGACAGGGGCAAGCAGTCCCTGGAGACAATCTGCCTCTT
GCTGGCCTACAAAATCAAGTATCCGGAGAACTTCTTTCTTCTCAGAGGGAACCACGAGTGCGCCAGCATCAATAGGATCT
ACGGATTTTATGATGAGTGTAAAAGAAGATACAACATTAAGCTGTGGAAAACGTTCACAGACTGTTTTAACTGCTTGCCG
ATAGCAGCCATCGTGGACGAGAAGATATTCTGCTGTCATGGAGGTTTATCACCAGATCTTCAATCTATGGAGCAGATTCG
GCGAATTATGAGACCAACTGATGTACCAGATCAAGGTCTTCTTTGTGATCTTTTGTGGTCTGACCCCGATAAAGATGTCT
TAGGCTGGGGTGAAAATGACAGAGGAGTGTCCTTCACATTTGGTGCAGAAGTGGTTGCAAAATTTCTCCATAAGCATGAT
TTGGATCTTATATGTAGAGCCCATCAGGTGGTTGAAGATGGCTATGAGTTTTTTGCAAAGAGGCAGTTAGTCACTCTGTT
TTCTGCACCCAACTACTGTGGCGAGTTTGACAATGCAGGCGCCATGATGAGTGTGGATGAGACCCTCATGTGTTCCTTCC
AGATTTTAAAGCCTGCAGAGAAAAAGAAGCCCAACGCCACGAGACCTGTCACACCACCACGGGGTATGATCACAAAGCAA
GCAAAGAAA
ATGGCGGATATCGACAAACTCAACATCGACAGCATCATCCAACGGCTGCTGGAAGTGAGAGGGTCCAAGCCAGGCAAGAA
TGTCCAGCTCCAGGAGAACGAGATCCGAGGACTCTGCCTGAAGTCTCGGGAGATCTTCCTCAGTCAGCCTATCCTTTTAG
AACTTGAAGCACCACTCAAGATATGTGGTGACATCCACGGGCAGTACTATGATTTGCTCCGTCTGTTTGAATACGGTGGC
TTTCCTCCAGAGAGCAACTATTTGTTTCTCGGGGACTATGTGGACAGGGGCAAGCAGTCCCTGGAGACAATCTGCCTCTT
GCTGGCCTACAAAATCAAGTATCCGGAGAACTTCTTTCTTCTCAGAGGGAACCACGAGTGCGCCAGCATCAATAGGATCT
ACGGATTTTATGATGAGTGTAAAAGAAGATACAACATTAAGCTGTGGAAAACGTTCACAGACTGTTTTAACTGCTTGCCG
ATAGCAGCCATCGTGGACGAGAAGATATTCTGCTGTCATGGAGGTTTATCACCAGATCTTCAATCTATGGAGCAGATTCG
GCGAATTATGAGACCAACTGATGTACCAGATCAAGGTCTTCTTTGTGATCTTTTGTGGTCTGACCCCGATAAAGATGTCT
TAGGCTGGGGTGAAAATGACAGAGGAGTGTCCTTCACATTTGGTGCAGAAGTGGTTGCAAAATTTCTCCATAAGCATGAT
TTGGATCTTATATGTAGAGCCCATCAGGTGGTTGAAGATGGCTATGAGTTTTTTGCAAAGAGGCAGTTAGTCACTCTGTT
TTCTGCACCCAACTACTGTGGCGAGTTTGACAATGCAGGCGCCATGATGAGTGTGGATGAGACCCTCATGTGTTCCTTCC
AGATTTTAAAGCCTGCAGAGAAAAAGAAGCCCAACGCCACGAGACCTGTCACACCACCACGGGGTATGATCACAAAGCAA
GCAAAGAAA
ORF - retro_mmus_2968 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 100. % |
| Parental protein coverage: | 100. % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | MADIDKLNIDSIIQRLLEVRGSKPGKNVQLQENEIRGLCLKSREIFLSQPILLELEAPLKICGDIHGQYY |
| MADIDKLNIDSIIQRLLEVRGSKPGKNVQLQENEIRGLCLKSREIFLSQPILLELEAPLKICGDIHGQYY | |
| Retrocopy | MADIDKLNIDSIIQRLLEVRGSKPGKNVQLQENEIRGLCLKSREIFLSQPILLELEAPLKICGDIHGQYY |
| Parental | DLLRLFEYGGFPPESNYLFLGDYVDRGKQSLETICLLLAYKIKYPENFFLLRGNHECASINRIYGFYDEC |
| DLLRLFEYGGFPPESNYLFLGDYVDRGKQSLETICLLLAYKIKYPENFFLLRGNHECASINRIYGFYDEC | |
| Retrocopy | DLLRLFEYGGFPPESNYLFLGDYVDRGKQSLETICLLLAYKIKYPENFFLLRGNHECASINRIYGFYDEC |
| Parental | KRRYNIKLWKTFTDCFNCLPIAAIVDEKIFCCHGGLSPDLQSMEQIRRIMRPTDVPDQGLLCDLLWSDPD |
| KRRYNIKLWKTFTDCFNCLPIAAIVDEKIFCCHGGLSPDLQSMEQIRRIMRPTDVPDQGLLCDLLWSDPD | |
| Retrocopy | KRRYNIKLWKTFTDCFNCLPIAAIVDEKIFCCHGGLSPDLQSMEQIRRIMRPTDVPDQGLLCDLLWSDPD |
| Parental | KDVLGWGENDRGVSFTFGAEVVAKFLHKHDLDLICRAHQVVEDGYEFFAKRQLVTLFSAPNYCGEFDNAG |
| KDVLGWGENDRGVSFTFGAEVVAKFLHKHDLDLICRAHQVVEDGYEFFAKRQLVTLFSAPNYCGEFDNAG | |
| Retrocopy | KDVLGWGENDRGVSFTFGAEVVAKFLHKHDLDLICRAHQVVEDGYEFFAKRQLVTLFSAPNYCGEFDNAG |
| Parental | AMMSVDETLMCSFQILKPAEKKKPNATRPVTPPRGMITKQAKK |
| AMMSVDETLMCSFQILKPAEKKKPNATRPVTPPRGMITKQAKK | |
| Retrocopy | AMMSVDETLMCSFQILKPAEKKKPNATRPVTPPRGMITKQAKK |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .54 RPM | 28 .99 RPM |
| SRP007412_cerebellum | 1 .09 RPM | 40 .08 RPM |
| SRP007412_heart | 0 .75 RPM | 17 .50 RPM |
| SRP007412_kidney | 1 .23 RPM | 24 .10 RPM |
| SRP007412_liver | 0 .27 RPM | 11 .96 RPM |
| SRP007412_testis | 7 .41 RPM | 147 .23 RPM |
RNA Polymerase II actvity near the 5' end of retro_mmus_2968 was not detected
16 EST(s) were mapped to retro_mmus_2968 retrocopy
| EST ID | Start | End | Identity | Match | Mis-match | Score |
|---|---|---|---|---|---|---|
| AL034693 | 119740470 | 119740753 | 100 | 282 | 0 | 281 |
| AV578978 | 119740657 | 119740879 | 100 | 222 | 0 | 222 |
| AW209791 | 119740111 | 119740571 | 99.8 | 458 | 0 | 455 |
| AW682422 | 119740330 | 119740714 | 100 | 384 | 0 | 384 |
| BG142781 | 119740186 | 119740571 | 100 | 385 | 0 | 385 |
| CA538375 | 119740622 | 119740820 | 100 | 198 | 0 | 198 |
| CD560144 | 119740047 | 119740570 | 100 | 523 | 0 | 523 |
| CF104161 | 119740104 | 119740571 | 99.2 | 463 | 4 | 459 |
| CF982022 | 119740505 | 119740740 | 100 | 235 | 0 | 235 |
| CJ047883 | 119740184 | 119740592 | 100 | 408 | 0 | 408 |
| CJ055725 | 119740340 | 119740723 | 99.5 | 381 | 2 | 379 |
| CJ137565 | 119740187 | 119740633 | 100 | 445 | 0 | 445 |
| CN707525 | 119740165 | 119740649 | 100 | 484 | 0 | 484 |
| CX233436 | 119740381 | 119740654 | 99.3 | 271 | 2 | 269 |
| CX234471 | 119740526 | 119740735 | 100 | 209 | 0 | 209 |
| DT911769 | 119740115 | 119740594 | 99.8 | 477 | 1 | 475 |
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_128187 | 222 libraries | 586 libraries | 259 libraries | 2 libraries | 3 libraries |
The graphical summary, for retro_mmus_2968 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
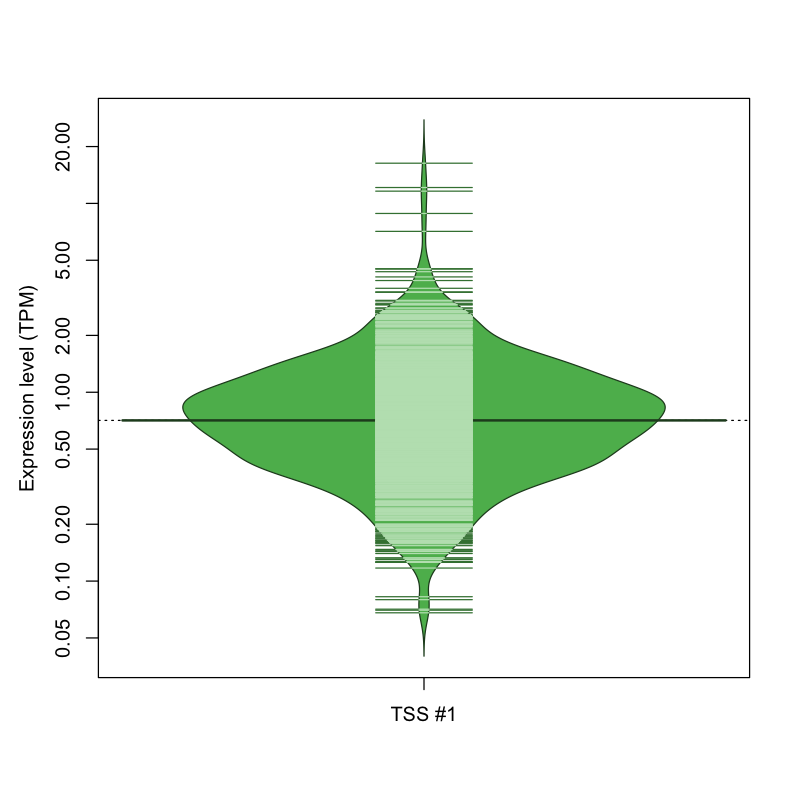
retro_mmus_2968 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_2968 has 0 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 10 parental genes, and 26 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Ailuropoda melanoleuca | ENSAMEG00000013861 | 1 retrocopy | |
| Bos taurus | ENSBTAG00000011198 | 2 retrocopies | |
| Ciona intestinalis | ENSCING00000004542 | 1 retrocopy | |
| Callithrix jacchus | ENSCJAG00000009147 | 7 retrocopies | |
| Equus caballus | ENSECAG00000017228 | 1 retrocopy | |
| Echinops telfairi | ENSETEG00000015721 | 1 retrocopy | |
| Myotis lucifugus | ENSMLUG00000010661 | 2 retrocopies | |
| Mus musculus | ENSMUSG00000004455 | 5 retrocopies | |
| Sorex araneus | ENSSARG00000000571 | 1 retrocopy | |
| Drosophila melanogaster | FBgn0003134 | 5 retrocopies |