RetrogeneDB ID: | retro_mmus_108 | ||
Retrocopylocation | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 12:73545277..73546339(-) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | ENSMUSG00000034435 | |
| Aliases: | Tmem30b, 9130011B11Rik | ||
| Status: | KNOWN_PROTEIN_CODING | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | Tmem30a | ||
| Ensembl ID: | ENSMUSG00000032328 | ||
| Aliases: | Tmem30a, 2010200I23Rik, AW540225, Cdc50a, D9Wsu20e | ||
| Description: | transmembrane protein 30A [Source:MGI Symbol;Acc:MGI:106402] |

Retrocopy-Parental alignment summary:
>retro_mmus_108
ATGACCTGGAGCGCCACGGCGCGGGGCGCGCACCAGCCCGACAACACCGCCTTCACTCAGCAGCGCCTCCCCGCCTGGCA
GCCGCTGCTGTCGGCGGGCATCGCGCTGCCGCTCTTCTTCTGCGCCGGCCTGGCGTTCATCGGCCTGGGCCTGGGCCTCT
TCTACTCCTCCAACGGCATCAAGGAGCTGGAGTACGACTACACCGGCAACCCCGGCACCGGCGATTGCTCGGTGTGCGCC
GCCAAGGGCCAGGGCCGCGCTCCGCCGCCCGGCTGCGCTTGCTCCTGGAGCTTTACGCTGCCCGAGCTCTTCCCGGGCCC
CGTGTACCTCTACTACGAGCTGTCCAACTTCTACCAGAACAACCGGCGCTACGGCGTGTCCCGCGACGACGCGCAGCTCA
GCGGCCTGGCCAGCGCGCTGCGCCACCCGGCCAACGAGTGCGCCCCCTACCAGTTCCGCTCCGACGGTCTGCCTATCGCA
CCGTGCGGCGCCATCGCCAACAGCCTCTTCAACGACTCCTTCTCGCTCTGGCACCAGCGCCAGCCCTCGGACCCCTTCGT
CGAGGTGCCGCTCGATCGCACCGCCATCGCCTGGTGGACCGACTACCACGTCAAGTTCCGCAACCCGCCGCTGGTGAACG
GCAGCCTGGCGCTGGCCTTCCGCGGCACGGCGCCGCCGCCCAACTGGCACCGGCCGGTTTATGAGCTCAGCCCGGATCCC
AACAACACCGGCTTCATCAACCAGGACTTCGTGGTGTGGATGCGCACCGCGGCGCTGCCCACGTTCCGCAAGCTCTATGC
GCGCATCCGCCAGGGCAACTACTCCGCCGGTCTGCCCCGGGGCACCTACCGTGTCAACATCACCTACAACTACCCGGTGC
GCGCCTTCGGCGGCCACAAGCTCATCATTCTTAGCAACATCTCTTGGATGGGTGGCAAGAACCCTTTCCTGGGCATCGCC
TACCTGGTCGTTGGCTCCCTCTGCATCGTCATGGGCTTTGTCATGCTGGTCGTCTACATTCGCTACCAGGACCAGGACGA
CGACGACAACGATGATGAGTGA
ATGACCTGGAGCGCCACGGCGCGGGGCGCGCACCAGCCCGACAACACCGCCTTCACTCAGCAGCGCCTCCCCGCCTGGCA
GCCGCTGCTGTCGGCGGGCATCGCGCTGCCGCTCTTCTTCTGCGCCGGCCTGGCGTTCATCGGCCTGGGCCTGGGCCTCT
TCTACTCCTCCAACGGCATCAAGGAGCTGGAGTACGACTACACCGGCAACCCCGGCACCGGCGATTGCTCGGTGTGCGCC
GCCAAGGGCCAGGGCCGCGCTCCGCCGCCCGGCTGCGCTTGCTCCTGGAGCTTTACGCTGCCCGAGCTCTTCCCGGGCCC
CGTGTACCTCTACTACGAGCTGTCCAACTTCTACCAGAACAACCGGCGCTACGGCGTGTCCCGCGACGACGCGCAGCTCA
GCGGCCTGGCCAGCGCGCTGCGCCACCCGGCCAACGAGTGCGCCCCCTACCAGTTCCGCTCCGACGGTCTGCCTATCGCA
CCGTGCGGCGCCATCGCCAACAGCCTCTTCAACGACTCCTTCTCGCTCTGGCACCAGCGCCAGCCCTCGGACCCCTTCGT
CGAGGTGCCGCTCGATCGCACCGCCATCGCCTGGTGGACCGACTACCACGTCAAGTTCCGCAACCCGCCGCTGGTGAACG
GCAGCCTGGCGCTGGCCTTCCGCGGCACGGCGCCGCCGCCCAACTGGCACCGGCCGGTTTATGAGCTCAGCCCGGATCCC
AACAACACCGGCTTCATCAACCAGGACTTCGTGGTGTGGATGCGCACCGCGGCGCTGCCCACGTTCCGCAAGCTCTATGC
GCGCATCCGCCAGGGCAACTACTCCGCCGGTCTGCCCCGGGGCACCTACCGTGTCAACATCACCTACAACTACCCGGTGC
GCGCCTTCGGCGGCCACAAGCTCATCATTCTTAGCAACATCTCTTGGATGGGTGGCAAGAACCCTTTCCTGGGCATCGCC
TACCTGGTCGTTGGCTCCCTCTGCATCGTCATGGGCTTTGTCATGCTGGTCGTCTACATTCGCTACCAGGACCAGGACGA
CGACGACAACGATGATGAGTGA
ORF - retro_mmus_108 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 57.14 % |
| Parental protein coverage: | 70.6 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | CACTINFTLKQSFEGNVFMYYGLSNFYQNHRRYVKSRDDSQLNGDPSALLNPSKECEPYR-RNEDRPIAP |
| CAC...FTL...F.G.V..YY.LSNFYQN.RRY..SRDD.QL.G..SAL..P..EC.PY..R....PIAP | |
| Retrocopy | CACSWSFTLPELFPGPVYLYYELSNFYQNNRRYGVSRDDAQLSGLASALRHPANECAPYQFRSDGLPIAP |
| Parental | CGAIANSMFNDTLELYLVANESDPKPIPIPLKKKGIAWWTDKNVKFRNPP-GKESLEEKFKDTIKPVNWH |
| CGAIANS.FND...L......SDP.....PL....IAWWTD..VKFRNPP....SL...F..T..P.NWH | |
| Retrocopy | CGAIANSLFNDSFSLWHQRQPSDPF-VEVPLDRTAIAWWTDYHVKFRNPPLVNGSLALAFRGTAPPPNWH |
| Parental | KAVYELDPEDESNNGFINEDFIVWMRTAALPTFRKLYRLIERRDDLHPTLPAGQYFLNITYNYPVHSFDG |
| ..VYEL.P.D..N.GFIN.DF.VWMRTAALPTFRKLY..I.R.......LP.G.Y..NITYNYPV..F.G | |
| Retrocopy | RPVYELSP-DPNNTGFINQDFVVWMRTAALPTFRKLYARI-RQGNYSAGLPRGTYRVNITYNYPVRAFGG |
| Parental | RKRMILSTISWMGGKNPFLGIAYITIGSISFLLGVVLLVINHKYRNSSN |
| .K..ILS.ISWMGGKNPFLGIAY...GS.....G.V.LV....Y..... | |
| Retrocopy | HKLIILSNISWMGGKNPFLGIAYLVVGSLCIVMGFVMLVVYIRYQDQDD |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .02 RPM | 303 .44 RPM |
| SRP007412_cerebellum | 0 .13 RPM | 193 .92 RPM |
| SRP007412_heart | 0 .09 RPM | 68 .74 RPM |
| SRP007412_kidney | 5 .22 RPM | 129 .84 RPM |
| SRP007412_liver | 8 .52 RPM | 138 .56 RPM |
| SRP007412_testis | 0 .05 RPM | 70 .30 RPM |
RNA Polymerase II actvity near the 5' end of retro_mmus_108 was not detected
No EST(s) were mapped for retro_mmus_108 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_24312 | 753 libraries | 187 libraries | 82 libraries | 36 libraries | 14 libraries |
| TSS #2 | TSS_24313 | 889 libraries | 139 libraries | 43 libraries | 1 library | 0 libraries |
| TSS #3 | TSS_24314 | 594 libraries | 122 libraries | 115 libraries | 63 libraries | 178 libraries |
| TSS #4 | TSS_24315 | 602 libraries | 111 libraries | 111 libraries | 49 libraries | 199 libraries |
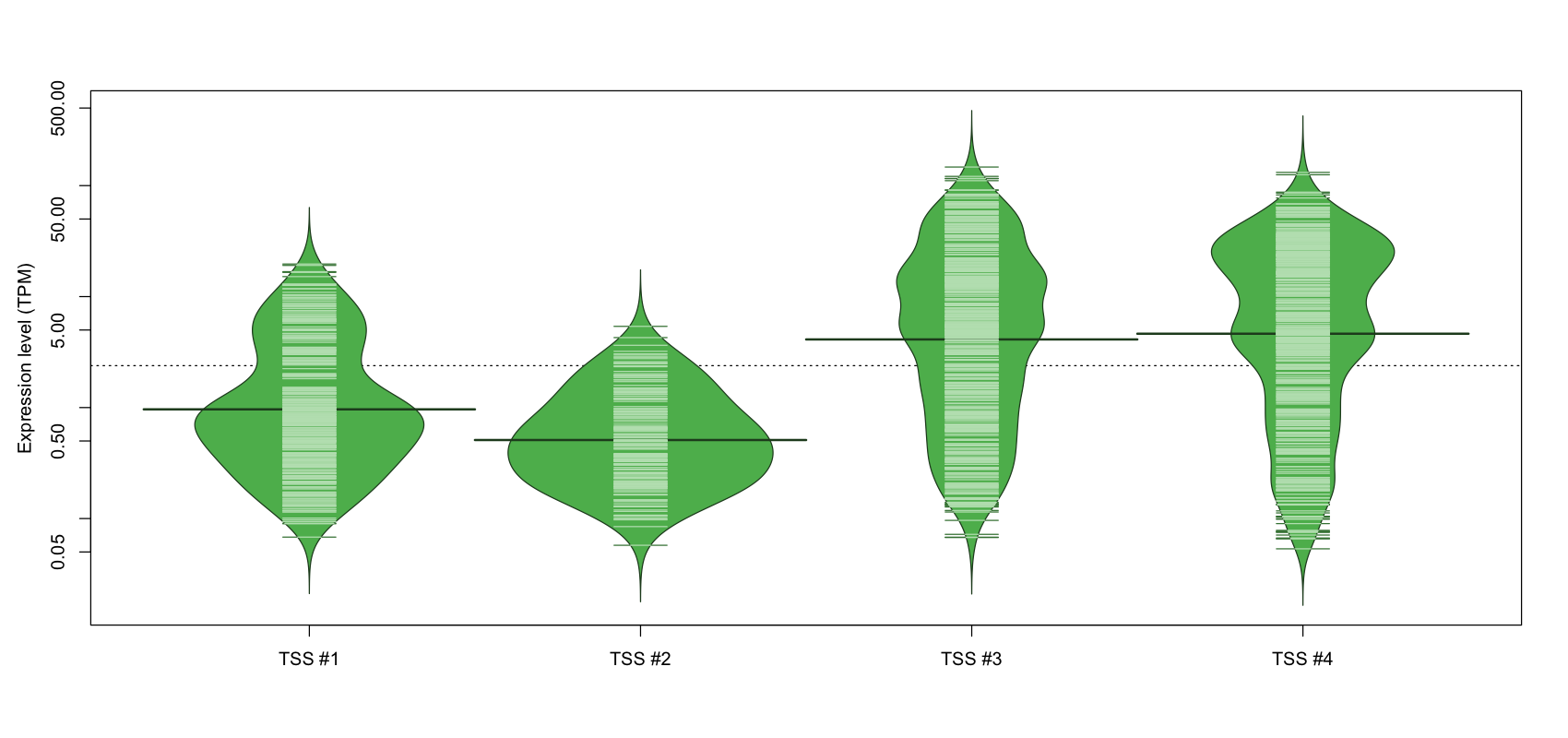
The graphical summary, for retro_mmus_108 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
retro_mmus_108 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_108 has 0 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 15 parental genes, and 15 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Bos taurus | ENSBTAG00000005100 | 1 retrocopy | |
| Canis familiaris | ENSCAFG00000002708 | 1 retrocopy | |
| Callithrix jacchus | ENSCJAG00000010325 | 1 retrocopy | |
| Cavia porcellus | ENSCPOG00000020712 | 1 retrocopy | |
| Homo sapiens | ENSG00000112697 | 1 retrocopy | |
| Myotis lucifugus | ENSMLUG00000011923 | 1 retrocopy | |
| Monodelphis domestica | ENSMODG00000018502 | 1 retrocopy | |
| Mus musculus | ENSMUSG00000032328 | 1 retrocopy |
retro_mmus_108 ,
|
| Nomascus leucogenys | ENSNLEG00000004962 | 1 retrocopy | |
| Otolemur garnettii | ENSOGAG00000015718 | 1 retrocopy | |
| Pongo abelii | ENSPPYG00000016782 | 1 retrocopy | |
| Pelodiscus sinensis | ENSPSIG00000007174 | 1 retrocopy | |
| Rattus norvegicus | ENSRNOG00000010895 | 1 retrocopy | |
| Ictidomys tridecemlineatus | ENSSTOG00000002543 | 1 retrocopy | |
| Tarsius syrichta | ENSTSYG00000008211 | 1 retrocopy |