RetrogeneDB ID: | retro_mmus_555 | ||
Retrocopy location | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 10:43319382..43319624(+) | ||
| Located in intron of: | ENSMUSG00000038240 | ||
Retrocopy information | Ensembl ID: | ENSMUSG00000089673 | |
| Aliases: | None | ||
| Status: | KNOWN_PSEUDOGENE | ||
Parental gene information | Parental gene summary: | ||
| Parental gene symbol: | Pole3 | ||
| Ensembl ID: | ENSMUSG00000028394 | ||
| Aliases: | None | ||
| Description: | polymerase (DNA directed), epsilon 3 (p17 subunit) [Source:MGI Symbol;Acc:MGI:1933378] |

Retrocopy-Parental alignment summary:
>retro_mmus_555
AAATCTGCCTGAAGCGCCATCTCCCAAGCCACCAGTGTCTTTGTTTTATATGCCACATCCTGTGCCAATAACTTCTCAAT
GAAAGGAAAGCACAAGACTGTGAATGCCAGTGATATGCCATCAGCCATGGAAGAGATGGAGTTCCAGCGGTTCCTAACAC
CTTGAAAGAAGCTCTAGAAGCACACAGGCGGGAGCAGAAAGGCAAGAAGGAGGCTTCAGAGCAGAAGAAGGACGACAAAG
AC
ORF - retro_mmus_555 Open Reading Frame is not conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 84.15 % |
| Parental protein coverage: | 55.86 % |
| Number of stop codons detected: | 1 |
| Number of frameshifts detected: | 1 |
Retrocopy - Parental Gene Alignment:
| Parental | KEARSAISRAASVFVLYATSCANNFAMKGKRKTLNASDVLSAMEEMEFQRFITP-LKEALEAYRREQKGK |
| K.A.SAIS.A.SVFVLYATSCANNF.MKGK.KT.NASD..SAMEEMEFQRF.TP.LKEALEA.RREQKGK | |
| Retrocopy | KSA*SAISQATSVFVLYATSCANNFSMKGKHKTVNASDMPSAMEEMEFQRFLTP<LKEALEAHRREQKGK |
| Parental | KEASEQKKKDKD |
| KEASEQKK.DKD | |
| Retrocopy | KEASEQKKDDKD |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .02 RPM | 19 .51 RPM |
| SRP007412_cerebellum | 0 .00 RPM | 29 .70 RPM |
| SRP007412_heart | 0 .00 RPM | 18 .77 RPM |
| SRP007412_kidney | 0 .00 RPM | 36 .15 RPM |
| SRP007412_liver | 0 .00 RPM | 21 .38 RPM |
| SRP007412_testis | 0 .05 RPM | 124 .23 RPM |
RNA Polymerase II activity near the 5' end of retro_mmus_555 was not detected
No EST(s) were mapped for retro_mmus_555 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_3388 | 454 libraries | 74 libraries | 472 libraries | 65 libraries | 7 libraries |
The graphical summary, for retro_mmus_555 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
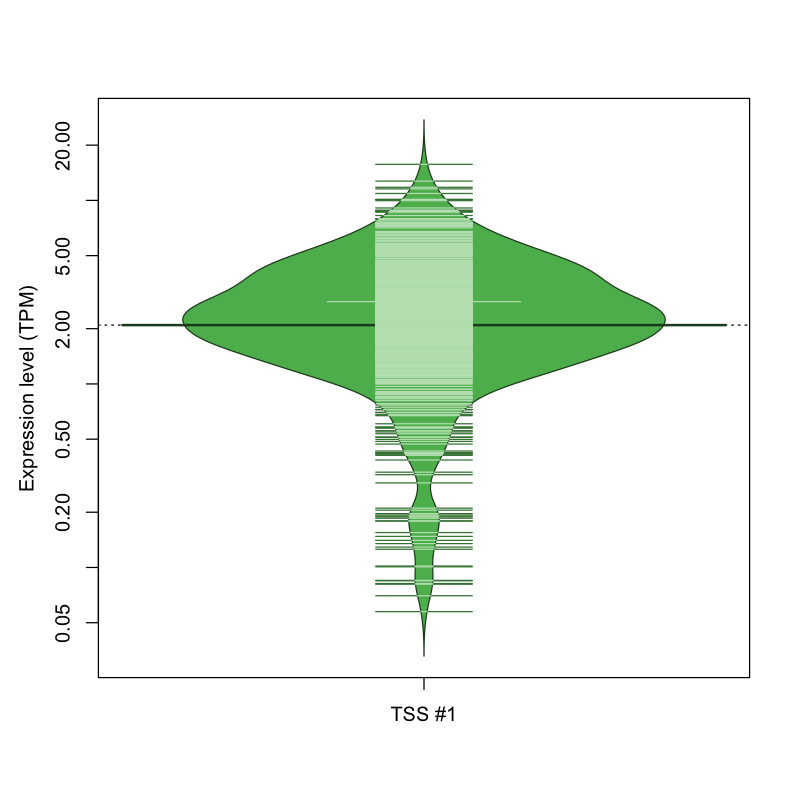
retro_mmus_555 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_555 has 0 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 9 parental genes, and 13 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Callithrix jacchus | ENSCJAG00000017917 | 1 retrocopy | |
| Cavia porcellus | ENSCPOG00000000313 | 1 retrocopy | |
| Dipodomys ordii | ENSDORG00000003073 | 2 retrocopies | |
| Monodelphis domestica | ENSMODG00000004358 | 3 retrocopies | |
| Mus musculus | ENSMUSG00000028394 | 2 retrocopies |
retro_mmus_124, retro_mmus_555 ,
|
| Pteropus vampyrus | ENSPVAG00000004717 | 1 retrocopy | |
| Rattus norvegicus | ENSRNOG00000015392 | 1 retrocopy | |
| Ictidomys tridecemlineatus | ENSSTOG00000027365 | 1 retrocopy | |
| Tupaia belangeri | ENSTBEG00000005211 | 1 retrocopy |