RetrogeneDB ID: | retro_mmus_32 | ||
Retrocopy location | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 6:48939788..48940127(-) | ||
| Located in intron of: | None | ||
Retrocopy information | Ensembl ID: | ENSMUSG00000068537 | |
| Aliases: | None | ||
| Status: | KNOWN_PROTEIN_CODING | ||
Parental gene information | Parental gene summary: | ||
| Parental gene symbol: | Cox7a2l | ||
| Ensembl ID: | ENSMUSG00000024248 | ||
| Aliases: | Cox7a2l, COX7AR, COX7RP, EB1, SIG-81, SIG81, Silg81 | ||
| Description: | cytochrome c oxidase subunit VIIa polypeptide 2-like [Source:MGI Symbol;Acc:MGI:106015] |

Retrocopy-Parental alignment summary:
>retro_mmus_32
GTGTACTATAAGTTTAGCAGGTCCACGCAGAAGTTGGCTAGAGCTTGGGCTTCGAAAGCCTATACTCCGCAGGGGTTACA
GCTGGTTTCCATGGAAACACCACCTACCACATTTGCTACACCAACCAAACTGACCTCCAGTGCAACAGCATATGATTATG
CTGGGAAGAACAAAGTTCCAGAGCTGCAGAAGTTCTTCCAGAAGGCTGATGGCATGCCCATTCACCTGAAAGGAGGCCTT
CCGGACCAAATGCTTTACCAGACCACCATGGCTCTGACAATGGGAGGGACCATCTACTGCCTGACTCTACATGGCCTTGC
AGCCCAGAAACTTGCTTGC
ORF - retro_mmus_32 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 77.06 % |
| Parental protein coverage: | 96.4 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected: | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | MYYKFSSFTQKLAGAWASEAYTPQGLKPVSTEAPPIIFATPTKLTSSVTAYDYSGKNKVPELQKFFQKAD |
| .YYKFS..TQKLA.AWAS.AYTPQGL..VS.E.PP..FATPTKLTSS.TAYDY.GKNKVPELQKFFQKAD | |
| Retrocopy | VYYKFSRSTQKLARAWASKAYTPQGLQLVSMETPPTTFATPTKLTSSATAYDYAGKNKVPELQKFFQKAD |
| Parental | G--FHLKRGLPDQMLYRTTMALTLGGTIYCLIALYMASQ |
| G...HLK.GLPDQMLY.TTMALT.GGTIYCL.....A.Q | |
| Retrocopy | GMPIHLKGGLPDQMLYQTTMALTMGGTIYCLTLHGLAAQ |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .00 RPM | 41 .17 RPM |
| SRP007412_cerebellum | 0 .00 RPM | 28 .01 RPM |
| SRP007412_heart | 0 .00 RPM | 67 .74 RPM |
| SRP007412_kidney | 0 .00 RPM | 44 .62 RPM |
| SRP007412_liver | 0 .00 RPM | 28 .76 RPM |
| SRP007412_testis | 0 .00 RPM | 22 .00 RPM |
RNA Polymerase II activity near the 5' end of retro_mmus_32 was not detected
No EST(s) were mapped for retro_mmus_32 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_122929 | 324 libraries | 580 libraries | 166 libraries | 0 libraries | 2 libraries |
The graphical summary, for retro_mmus_32 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
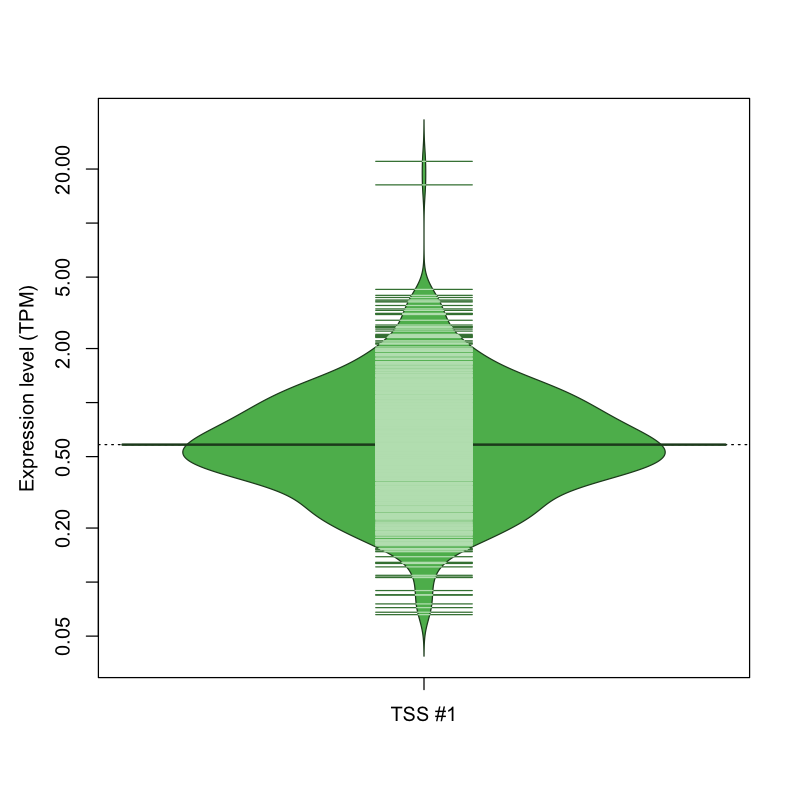
retro_mmus_32 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_32 has 0 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 15 parental genes, and 20 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Canis familiaris | ENSCAFG00000032040 | 1 retrocopy | |
| Callithrix jacchus | ENSCJAG00000014705 | 1 retrocopy | |
| Echinops telfairi | ENSETEG00000011940 | 1 retrocopy | |
| Homo sapiens | ENSG00000115944 | 1 retrocopy | |
| Gorilla gorilla | ENSGGOG00000010739 | 1 retrocopy | |
| Macaca mulatta | ENSMMUG00000016278 | 1 retrocopy | |
| Mustela putorius furo | ENSMPUG00000008875 | 1 retrocopy | |
| Mus musculus | ENSMUSG00000024248 | 2 retrocopies |
retro_mmus_32 , retro_mmus_926,
|
| Nomascus leucogenys | ENSNLEG00000016666 | 1 retrocopy | |
| Oryctolagus cuniculus | ENSOCUG00000027189 | 3 retrocopies | |
| Otolemur garnettii | ENSOGAG00000016029 | 1 retrocopy | |
| Pan troglodytes | ENSPTRG00000011865 | 1 retrocopy | |
| Sarcophilus harrisii | ENSSHAG00000012014 | 1 retrocopy | |
| Ictidomys tridecemlineatus | ENSSTOG00000012922 | 1 retrocopy | |
| Tupaia belangeri | ENSTBEG00000014124 | 3 retrocopies |