RetrogeneDB ID: | retro_mmus_14 | ||
Retrocopy location | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 12:119260977..119262855(+) | ||
| Located in intron of: | None | ||
Retrocopy information | Ensembl ID: | ENSMUSG00000021908 | |
| Aliases: | None | ||
| Status: | KNOWN_PROTEIN_CODING | ||
Parental gene information | Parental gene summary: | ||
| Parental gene symbol: | Ncoa4 | ||
| Ensembl ID: | ENSMUSG00000056234 | ||
| Aliases: | Ncoa4, ARA70, Rfg | ||
| Description: | nuclear receptor coactivator 4 [Source:MGI Symbol;Acc:MGI:1350932] |

Retrocopy-Parental alignment summary:
>retro_mmus_14
ATGAACACATCCCTGGAACAGAGTGGCTGCTACAGTAATAGAGAAACTCTTCTGAGGTGCAGTGATGCAAGGAGGGAGCT
GGAGCTTGCTATTGGTGGAGTACTTCGGGCTGAACAGCAAATTAAAGATAATTTACGAGAGGTAAAAGCTCAGATTCACA
GCTGCATAAGTCGCCACCTGGAATGCCTTCGGAGCCGAGAGGTGTGGCTCAATGAACAGGTCGATCTCATCTATCAGCTT
AAAGAGGAGACACTGCAGCAGCAGGCTCAGCAGCTCTATTGGCTAATGGGCCAATTCAATTGTCTTATTCATCAACTGGA
GTATACTCAGAACAAAGATCTTGCCAATCAAGTCTCTGTGTGCTTGGAGAGACTGGGCAGTTTGGCCCTGAAGCCTGAAG
ATTCAACTGTCCTACTCTTTGAAGCAGATACATCTGCTCTGCGCCAGACCATCACCACATTTGGATCCCTTAAGACCATT
CAAATCCCTGAGCATCTGATGGCTCACGCGAGCTCCTCAAGTATTGGGCCTTTCCTAGAGAAAAGAGGCTATATCCAGGT
GCCAGAGCAGAAGTCAGCATCCAGTGGCACAGCTGTATCTCTGAGTGAGTGGCTTCTTGTAAGCAAACCTGCCATTGGTC
TTCAGGCTCCTTATGTACCCAGTACCAACCCACAGGACTGGCTTATCCCAAAACAGACTTCAGAGAATAGTCAGACTTCT
GCCAGAGCCTGCAGTTTCTTCAGCGATGCCTGGGGCAATCTGAAGGGCTTAGAAAATTGGCTCCTTAACAGTCATCAACA
AGAAATTGCTGGAAAACCAAGTTCCTCAAAATGTAACAGCCATTGTAGCACCAGTTCTTTCTCTCCTGAAGCTGAAAAGG
CTGAAGATGTAGAGCTCCTTGATCAAGACGAGCTGGACCTGTCTGATTGGTTGGTGACTCCTCAGGAACCCTGTGAGCTG
GAGAAGCCTGTGGATGGCAGCTGGGAAACCAGCGAGAAGTTTAAACTCTTGTTCCAGGTGTTCAGAGAGCCCTACAATGT
GAGTGATTGGCTTGTCAAGCCTGACTCCTGTACCAACTGTCAGGGCAACCAGCCTAGAGGTGTGGAGATTGAGAACCTGG
GCAATCTGAAATGCCTAAATGACCACTTGGAGGCCAAAAAATCAGTGTCGGTCCCCGGCGCAACAATCAGCGACGGCTGG
CTTGCCCAGAACCATCAGGACACATGGAAAGTAGAGGAAGTGTGTAAAGCCAACGAGCCCTGCACCAGTTTTGCAGAGTG
TGTGTGTGATGACAACTGTGAGAAGGAAGCCGTGTATAAATGGCTTCTGAAGAAGGGAGGAAAGGACAAGAATGGAATGC
CTATGGAACCTAAATCTGAGCCTGAGAAACACAGAGAGTCCCTGACACTGTGGCTCTGCCCTTCCAGAAATGAGCTAACA
GAACAAGCTAAGGCACCCAAGGCTATGGCTCCTGCTAGAATTGCTGATTCCTTCCACGTCATAAAGAATAGTTCCTTGTC
AGAGTGGCTTATAGGGCCTACGTGCAAAGGAGGCCCTAAGGACGTGCCTAATACAGAGGAGAGAGCTGGCAAAGAGATGC
TTCAAAGCTCCATGGCCACTTCCTGGTGTCCCTTTAACACTGCCGACTGGGTTTTACCAGGAAAGAAAGTGGGAAGCCTC
AGCCAGTTCCCCTCTGGAGAAGATAAGTGGCTACTTCGAAAGAAGGCACAGGAAGCATTTCTTAATTCACCCCTACAAGA
AGAACGTAACTTTCGCCCCGACTGTTACGGCCTCCCAGCAGTTTGTGATCTCTTTGCCTGTATGCAGCTTAAGGTTGATA
AAGAAAAATGGCTGTACAGAACTCCTCTACAGATGTGA
ORF - retro_mmus_14 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 99.68 % |
| Parental protein coverage: | 100.0 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected: | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | MNTSLEQSGCYSNRETLLRCSDARRELELAIGGVLRAEQQIKDNLREVKAQIHSCISRHLECLRSREVWL |
| MNTSLEQSGCYSNRETLLRCSDARRELELAIGGVLRAEQQIKDNLREVKAQIHSCISRHLECLRSREVWL | |
| Retrocopy | MNTSLEQSGCYSNRETLLRCSDARRELELAIGGVLRAEQQIKDNLREVKAQIHSCISRHLECLRSREVWL |
| Parental | NEQVDLIYQLKEETLQQQAQQLYWLMGQFNCLIHQLEYTQNKDLANQVSVCLERLGSLALKPEDSTVLLF |
| NEQVDLIYQLKEETLQQQAQQLYWLMGQFNCLIHQLEYTQNKDLANQVSVCLERLGSLALKPEDSTVLLF | |
| Retrocopy | NEQVDLIYQLKEETLQQQAQQLYWLMGQFNCLIHQLEYTQNKDLANQVSVCLERLGSLALKPEDSTVLLF |
| Parental | EADTSALRQTITTFGSLKTIQIPEHLMAHASSSSIGPFLEKRGYIQVPEQKSASSGTAVSLSEWLLVSKP |
| EADTSALRQTITTFGSLKTIQIPEHLMAHASSSSIGPFLEKRGYIQVPEQKSASSGTAVSLSEWLLVSKP | |
| Retrocopy | EADTSALRQTITTFGSLKTIQIPEHLMAHASSSSIGPFLEKRGYIQVPEQKSASSGTAVSLSEWLLVSKP |
| Parental | AIGLQAPYVPSTNPQDWLIPKQTSENSQTSARACSFFSDAWGNLKGLENWLLNSHQQEIAGKPSSSKCNS |
| AIGLQAPYVPSTNPQDWLIPKQTSENSQTSARACSFFSDAWGNLKGLENWLLNSHQQEIAGKPSSSKCNS | |
| Retrocopy | AIGLQAPYVPSTNPQDWLIPKQTSENSQTSARACSFFSDAWGNLKGLENWLLNSHQQEIAGKPSSSKCNS |
| Parental | HCSTSSFSPEAEKAEDVELLDQDELDLSDWLVTPQEPCELEKPVDGSWETSEKFKLLFQVFREPYNVSDW |
| HCSTSSFSPEAEKAEDVELLDQDELDLSDWLVTPQEPCELEKPVDGSWETSEKFKLLFQVFREPYNVSDW | |
| Retrocopy | HCSTSSFSPEAEKAEDVELLDQDELDLSDWLVTPQEPCELEKPVDGSWETSEKFKLLFQVFREPYNVSDW |
| Parental | LVKPDSCTNCQGNQPRGVEIENLGNLKCLNDHLEAKKSVSVPGATISDGWLAQNHQDTWKVEEVCKANEP |
| LVKPDSCTNCQGNQPRGVEIENLGNLKCLNDHLEAKKSVSVPGATISDGWLAQNHQDTWKVEEVCKANEP | |
| Retrocopy | LVKPDSCTNCQGNQPRGVEIENLGNLKCLNDHLEAKKSVSVPGATISDGWLAQNHQDTWKVEEVCKANEP |
| Parental | CTSFAECVCDDNCEKEAMYKWLLKKGGKDKNGMPMEPKSEPEKHRESLTLWLCPSRNELTEQAKAPKAMA |
| CTSFAECVCDDNCEKEA.YKWLLKKGGKDKNGMPMEPKSEPEKHRESLTLWLCPSRNELTEQAKAPKAMA | |
| Retrocopy | CTSFAECVCDDNCEKEAVYKWLLKKGGKDKNGMPMEPKSEPEKHRESLTLWLCPSRNELTEQAKAPKAMA |
| Parental | PARIADSFHVIKNSSLSEWLMGPTCKGGPKDVPNTEERAGKEMLQSSMATSWCPFNTADWVLPGKKVGSL |
| PARIADSFHVIKNSSLSEWL.GPTCKGGPKDVPNTEERAGKEMLQSSMATSWCPFNTADWVLPGKKVGSL | |
| Retrocopy | PARIADSFHVIKNSSLSEWLIGPTCKGGPKDVPNTEERAGKEMLQSSMATSWCPFNTADWVLPGKKVGSL |
| Parental | SQFPSGEDKWLLRKKAQEAFLNSPLQEERNFRPDCYGLPAVCDLFACMQLKVDKEKWLYRTPLQM |
| SQFPSGEDKWLLRKKAQEAFLNSPLQEERNFRPDCYGLPAVCDLFACMQLKVDKEKWLYRTPLQM | |
| Retrocopy | SQFPSGEDKWLLRKKAQEAFLNSPLQEERNFRPDCYGLPAVCDLFACMQLKVDKEKWLYRTPLQM |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .35 RPM | 15 .70 RPM |
| SRP007412_cerebellum | 1 .35 RPM | 11 .68 RPM |
| SRP007412_heart | 0 .31 RPM | 14 .41 RPM |
| SRP007412_kidney | 4 .20 RPM | 24 .27 RPM |
| SRP007412_liver | 3 .36 RPM | 29 .52 RPM |
| SRP007412_testis | 5 .17 RPM | 46 .93 RPM |
RNA Polymerase II activity near the 5' end of retro_mmus_14 was not detected
82 EST(s) were mapped to retro_mmus_14 retrocopy
| EST ID | Start | End | Identity | Match | Mis-match | Score |
|---|---|---|---|---|---|---|
| AA871762 | 119260944 | 119261595 | 100 | 651 | 0 | 651 |
| AI788032 | 119260933 | 119261520 | 98.2 | 573 | 10 | 561 |
| AI791039 | 119260933 | 119261562 | 99.6 | 624 | 3 | 621 |
| AI931301 | 119260933 | 119261460 | 99.7 | 525 | 2 | 523 |
| AI119109 | 119260933 | 119261464 | 100 | 529 | 0 | 529 |
| AV483501 | 119260962 | 119261442 | 100 | 480 | 0 | 480 |
| AV489667 | 119260955 | 119261455 | 99.4 | 494 | 2 | 490 |
| AW681837 | 119261638 | 119262051 | 99.8 | 409 | 1 | 408 |
| AW761850 | 119260979 | 119261556 | 99.7 | 574 | 2 | 572 |
| AW822645 | 119260962 | 119261541 | 99.4 | 575 | 4 | 571 |
| BG060958 | 119261271 | 119261660 | 99.8 | 388 | 1 | 387 |
| BG173428 | 119261143 | 119261867 | 98.2 | 690 | 6 | 668 |
| BM963299 | 119261305 | 119262049 | 99.9 | 734 | 1 | 725 |
| BQ179847 | 119261399 | 119262091 | 99.3 | 684 | 4 | 678 |
| BQ443829 | 119261587 | 119262279 | 99.9 | 690 | 1 | 689 |
| BY044839 | 119260932 | 119261309 | 99.8 | 376 | 1 | 375 |
| BY046307 | 119260935 | 119261310 | 99.8 | 374 | 1 | 373 |
| BY047079 | 119260935 | 119261328 | 99.8 | 392 | 1 | 391 |
| BY047296 | 119260932 | 119261318 | 100 | 385 | 0 | 385 |
| BY048150 | 119260935 | 119261339 | 98.8 | 398 | 5 | 393 |
| BY051809 | 119260932 | 119261304 | 100 | 372 | 0 | 372 |
| BY058824 | 119260935 | 119261329 | 100 | 394 | 0 | 394 |
| BY064014 | 119260931 | 119261310 | 99.8 | 378 | 1 | 377 |
| BY072446 | 119260936 | 119261313 | 99.8 | 376 | 1 | 375 |
| BY081426 | 119261119 | 119261470 | 99.2 | 346 | 3 | 343 |
| BY089934 | 119261117 | 119261490 | 100 | 371 | 0 | 371 |
| BY106212 | 119261071 | 119261411 | 100 | 340 | 0 | 340 |
| BY226295 | 119260935 | 119261352 | 100 | 417 | 0 | 417 |
| BY226675 | 119260935 | 119261348 | 100 | 413 | 0 | 413 |
| BY713890 | 119260931 | 119261931 | 98.5 | 943 | 8 | 914 |
| BY738478 | 119260932 | 119261589 | 100 | 656 | 0 | 656 |
| BY740613 | 119260931 | 119261560 | 99.1 | 616 | 6 | 610 |
| CA530233 | 119261271 | 119261874 | 100 | 600 | 0 | 599 |
| CA532563 | 119261788 | 119262361 | 99.9 | 572 | 1 | 571 |
| CA538207 | 119261057 | 119261417 | 100 | 360 | 0 | 360 |
| CA558474 | 119261464 | 119261976 | 99.9 | 511 | 1 | 510 |
| CD547859 | 119260960 | 119261500 | 100 | 540 | 0 | 540 |
| CD553975 | 119260962 | 119261491 | 100 | 529 | 0 | 529 |
| CD565398 | 119261081 | 119261508 | 100 | 427 | 0 | 427 |
| CF163860 | 119260944 | 119261556 | 99.9 | 611 | 1 | 610 |
| CF166272 | 119260956 | 119261424 | 99.6 | 466 | 2 | 464 |
| CF173561 | 119261464 | 119262120 | 100 | 656 | 0 | 656 |
| CF173948 | 119261309 | 119261938 | 100 | 629 | 0 | 629 |
| CF174176 | 119260972 | 119261658 | 99.9 | 685 | 1 | 684 |
| CF736162 | 119261191 | 119261863 | 99.9 | 669 | 1 | 668 |
| CF739905 | 119261302 | 119262050 | 100 | 744 | 0 | 741 |
| CF897386 | 119261209 | 119261827 | 99.9 | 617 | 1 | 616 |
| CF910282 | 119261001 | 119261598 | 100 | 597 | 0 | 597 |
| CJ093868 | 119260935 | 119261614 | 99.8 | 675 | 2 | 672 |
| CJ164328 | 119260931 | 119261375 | 100 | 444 | 0 | 444 |
| CK389649 | 119260950 | 119261308 | 100 | 358 | 0 | 358 |
| CK790719 | 119260962 | 119261614 | 100 | 645 | 0 | 638 |
| CN459674 | 119260944 | 119261720 | 100 | 774 | 0 | 773 |
| CN527720 | 119260962 | 119261690 | 100 | 725 | 0 | 724 |
| CN530256 | 119261019 | 119261828 | 100 | 800 | 0 | 794 |
| CN538994 | 119260962 | 119261777 | 100 | 812 | 0 | 811 |
| CN664047 | 119260958 | 119261589 | 100 | 631 | 0 | 631 |
| CN664751 | 119260944 | 119261540 | 100 | 596 | 0 | 596 |
| CN666978 | 119260944 | 119261500 | 100 | 556 | 0 | 556 |
| CN669808 | 119261351 | 119261938 | 100 | 587 | 0 | 587 |
| CN673558 | 119261373 | 119261904 | 99.9 | 530 | 1 | 529 |
| CN675041 | 119260988 | 119261540 | 100 | 552 | 0 | 552 |
| CN678529 | 119260949 | 119261508 | 100 | 559 | 0 | 559 |
| CN679714 | 119261202 | 119261747 | 100 | 545 | 0 | 545 |
| CN680754 | 119260976 | 119261446 | 99.8 | 469 | 1 | 468 |
| CN686961 | 119260941 | 119261500 | 100 | 559 | 0 | 559 |
| CN691660 | 119261017 | 119261595 | 100 | 578 | 0 | 578 |
| CN691798 | 119260962 | 119261352 | 100 | 390 | 0 | 390 |
| CN692685 | 119261132 | 119261483 | 99.8 | 350 | 1 | 349 |
| CN695594 | 119260944 | 119261366 | 100 | 422 | 0 | 422 |
| CN695978 | 119261410 | 119261903 | 100 | 493 | 0 | 493 |
| CN700241 | 119260980 | 119261542 | 99.9 | 561 | 1 | 560 |
| CN702075 | 119260950 | 119261519 | 100 | 569 | 0 | 569 |
| CN704921 | 119260980 | 119261402 | 99.8 | 421 | 1 | 420 |
| CN704992 | 119261318 | 119261747 | 100 | 429 | 0 | 429 |
| CR758207 | 119261076 | 119261636 | 99 | 554 | 6 | 548 |
| CV562600 | 119261156 | 119261921 | 99.8 | 761 | 1 | 757 |
| CV783866 | 119261202 | 119261994 | 100 | 788 | 0 | 787 |
| CX240505 | 119260962 | 119261743 | 100 | 781 | 0 | 781 |
| CX563420 | 119261292 | 119262154 | 99.9 | 857 | 1 | 855 |
| DT908197 | 119261078 | 119261633 | 99.5 | 552 | 3 | 549 |
| DT930084 | 119261085 | 119261633 | 100 | 545 | 0 | 542 |
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_22166 | 140 libraries | 167 libraries | 212 libraries | 125 libraries | 428 libraries |
The graphical summary, for retro_mmus_14 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
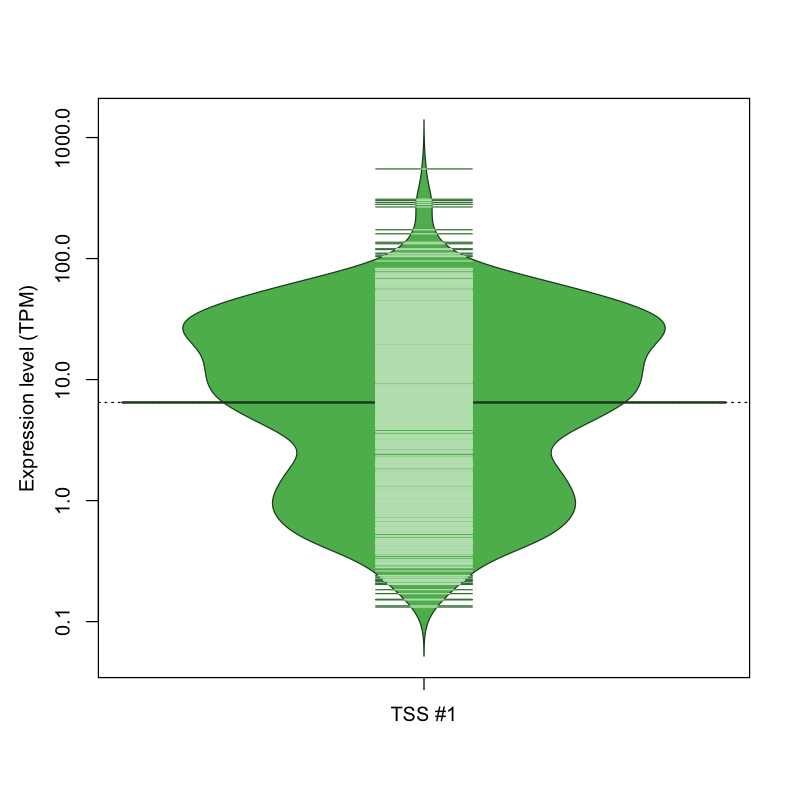
retro_mmus_14 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_14 has 0 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 5 parental genes, and 5 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Bos taurus | ENSBTAG00000019565 | 1 retrocopy | |
| Choloepus hoffmanni | ENSCHOG00000004343 | 1 retrocopy | |
| Callithrix jacchus | ENSCJAG00000013352 | 1 retrocopy | |
| Myotis lucifugus | ENSMLUG00000000619 | 1 retrocopy | |
| Mus musculus | ENSMUSG00000056234 | 1 retrocopy |
retro_mmus_14 ,
|