RetrogeneDB ID: | retro_mmus_2679 | ||
Retrocopylocation | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 5:104988979..104989329(-) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | None | |
| Aliases: | None | ||
| Status: | NOVEL | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | Sec61b | ||
| Ensembl ID: | ENSMUSG00000053317 | ||
| Aliases: | Sec61b, 1190006C12Rik, AI326121, AW122942 | ||
| Description: | Sec61 beta subunit [Source:MGI Symbol;Acc:MGI:1913462] |

Retrocopy-Parental alignment summary:
>retro_mmus_2679
GCCGGGTCCAAAGCCCAGTGGCACCAACGTGGGTTCCTCTGGCCACTCTCCCAGCAAAGCGGTGGCCACAAGGGCTGCAG
GATCCACTGTCTGGCAGAGAAAAAATGCCAGCTGTGGGACCCGGAGCGCAGGCCGCACCACCTCTGCAGGGACTGGGGGG
ATGTGGCGATTCTACACGGAAGATTCCTCAGGGCTCAAAGTGGGCCTTGTCCCAGTGCTGGTGATGAGTCTTCTGTTCAT
CGCTTCTGTATTTATGCTGAACATTTGGGGCAAGTACACGTGATCTTAGATTGGGCTACATCCATCTGTCATCTGAAGAA
GAAGAAGGAAAAAACCCAACATATCTTGGA
GCCGGGTCCAAAGCCCAGTGGCACCAACGTGGGTTCCTCTGGCCACTCTCCCAGCAAAGCGGTGGCCACAAGGGCTGCAG
GATCCACTGTCTGGCAGAGAAAAAATGCCAGCTGTGGGACCCGGAGCGCAGGCCGCACCACCTCTGCAGGGACTGGGGGG
ATGTGGCGATTCTACACGGAAGATTCCTCAGGGCTCAAAGTGGGCCTTGTCCCAGTGCTGGTGATGAGTCTTCTGTTCAT
CGCTTCTGTATTTATGCTGAACATTTGGGGCAAGTACACGTGATCTTAGATTGGGCTACATCCATCTGTCATCTGAAGAA
GAAGAAGGAAAAAACCCAACATATCTTGGA
ORF - retro_mmus_2679 Open Reading Frame is not conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 89.08 % |
| Parental protein coverage: | 84.29 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected | 1 |
Retrocopy - Parental Gene Alignment:
| Parental | AGSNAQWHQRGLLWPLSQQSGGRTGCGIHCSAEKKCQLWDPERRPHHLCRDWGDVAILHGRFPRAQSGPC |
| AGS.AQWHQRG.LWPLSQQSGG..GC.IHC.AEKKCQLWDPERRPHHLCRDWGDVAILHGRF.RAQSGPC | |
| Retrocopy | AGSKAQWHQRGFLWPLSQQSGGHKGCRIHCLAEKKCQLWDPERRPHHLCRDWGDVAILHGRFLRAQSGPC |
| Parental | PSAGDESSVHRCCIYAAHLGQVHAIIDWATSICHLKKKKKEK-NPTYLG |
| PSAGDESSVHR.CIYA.HLGQVH.I.DWATSICHL.KKKKEK.NPTYLG | |
| Retrocopy | PSAGDESSVHRFCIYAEHLGQVHVILDWATSICHL-KKKKEK<NPTYLG |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .02 RPM | 6 .59 RPM |
| SRP007412_cerebellum | 0 .00 RPM | 4 .69 RPM |
| SRP007412_heart | 0 .00 RPM | 4 .51 RPM |
| SRP007412_kidney | 0 .02 RPM | 10 .78 RPM |
| SRP007412_liver | 0 .00 RPM | 25 .67 RPM |
| SRP007412_testis | 0 .00 RPM | 6 .22 RPM |
RNA Polymerase II actvity near the 5' end of retro_mmus_2679 was not detected
No EST(s) were mapped for retro_mmus_2679 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_107586 | 26 libraries | 36 libraries | 208 libraries | 249 libraries | 553 libraries |
The graphical summary, for retro_mmus_2679 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
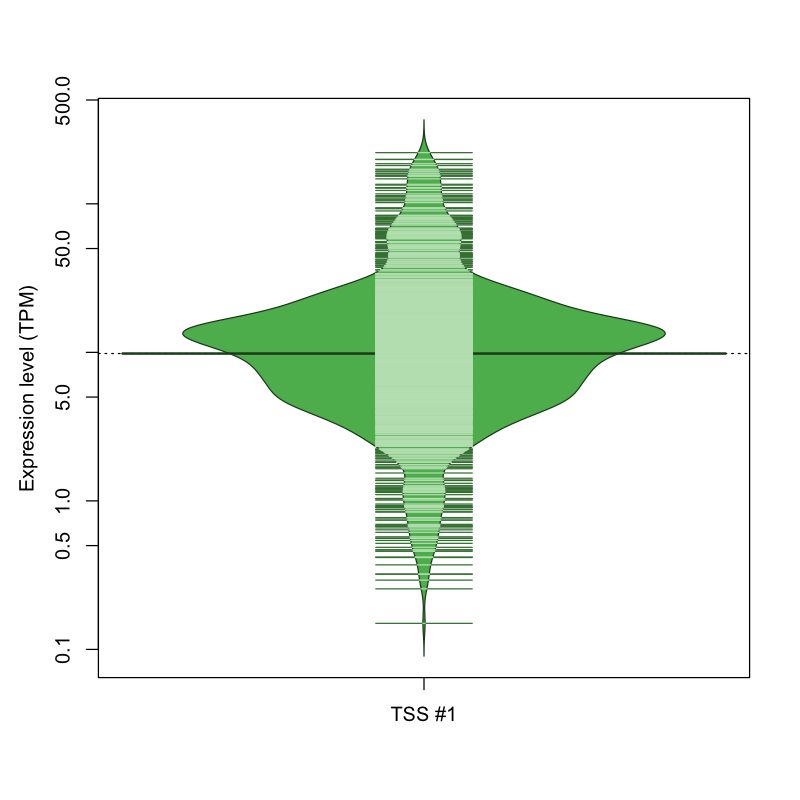
retro_mmus_2679 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_2679 has 0 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 16 parental genes, and 27 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Bos taurus | ENSBTAG00000002457 | 3 retrocopies | |
| Callithrix jacchus | ENSCJAG00000011279 | 1 retrocopy | |
| Equus caballus | ENSECAG00000026930 | 1 retrocopy | |
| Echinops telfairi | ENSETEG00000005293 | 4 retrocopies | |
| Homo sapiens | ENSG00000106803 | 1 retrocopy | |
| Gorilla gorilla | ENSGGOG00000004205 | 1 retrocopy | |
| Macropus eugenii | ENSMEUG00000000321 | 1 retrocopy | |
| Microcebus murinus | ENSMICG00000011121 | 4 retrocopies | |
| Myotis lucifugus | ENSMLUG00000001097 | 1 retrocopy | |
| Macaca mulatta | ENSMMUG00000010661 | 1 retrocopy | |
| Mus musculus | ENSMUSG00000053317 | 2 retrocopies |
retro_mmus_1377, retro_mmus_2679 ,
|
| Nomascus leucogenys | ENSNLEG00000017055 | 1 retrocopy | |
| Otolemur garnettii | ENSOGAG00000004168 | 3 retrocopies | |
| Pongo abelii | ENSPPYG00000019441 | 1 retrocopy | |
| Pan troglodytes | ENSPTRG00000021184 | 1 retrocopy | |
| Rattus norvegicus | ENSRNOG00000006345 | 1 retrocopy |