RetrogeneDB ID: | retro_mmus_244 | ||
Retrocopylocation | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 3:39358257..39358905(-) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | ENSMUSG00000050533 | |
| Aliases: | None | ||
| Status: | KNOWN_PROTEIN_CODING | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | Cflar | ||
| Ensembl ID: | ENSMUSG00000026031 | ||
| Aliases: | Cflar, 2310024N18Rik, A430105C05Rik, CLARP, Cash, Casper, ENSMUSG00000072980, FLAME, FLAME-1, Flip, Gm9845, I-FLICE, MRIT, c-Flip | ||
| Description: | CASP8 and FADD-like apoptosis regulator [Source:MGI Symbol;Acc:MGI:1336166] |

Retrocopy-Parental alignment summary:
>retro_mmus_244
ATGGCCCAGAGCCCTGTGTCTGCCGAGGTCATTCACCAGGTGGAAGAGTGTCTTGATGAAGACGAGAAGGAGATGATGCT
CTTCCTGTGTAGAGATGTGACTGAGAACCTGGCTGCACCTAACGTCAGGGACCTCCTGGATAGCTTAAGTGAGAGAGGCC
AGCTCTCTTTTGCTACCTTGGCTGAATTGCTCTACAGAGTGAGGCGGTTTGACCTTCTCAAGAGGATCTTGAAGACAGAC
AAAGCAACCGTGGAGGACCACCTGCGCAGAAACCCTCACCTGGTTTCTGATTATAGGGTCCTGCTGATGGAGATTGGTGA
GAGCTTAGATCAGAACGATGTATCCTCCTTAGTTTTCCTTACAAGGGATTACACAGGCAGAGGCAAGATAGCCAAGGACA
AGAGTTTCTTGGATCTGGTGATTGAATTGGAGAAACTGAATCTAATTGCTTCAGACCAATTGAATTTGTTAGAAAAATGC
CTGAAGAACATCCACAGAATAGACTTGAACACAAAGATCCAGAAGTACACCCAGTCCAGCCAAGGAGCAAGATCAAATAT
GAATACTCTCCAGGCTTCGCTCCCAAAATTGAGTATCAAGTATAACTCAAGGGTGAGTCTGGAGCCAGTGTATGGAGTAC
CAGCATGA
ATGGCCCAGAGCCCTGTGTCTGCCGAGGTCATTCACCAGGTGGAAGAGTGTCTTGATGAAGACGAGAAGGAGATGATGCT
CTTCCTGTGTAGAGATGTGACTGAGAACCTGGCTGCACCTAACGTCAGGGACCTCCTGGATAGCTTAAGTGAGAGAGGCC
AGCTCTCTTTTGCTACCTTGGCTGAATTGCTCTACAGAGTGAGGCGGTTTGACCTTCTCAAGAGGATCTTGAAGACAGAC
AAAGCAACCGTGGAGGACCACCTGCGCAGAAACCCTCACCTGGTTTCTGATTATAGGGTCCTGCTGATGGAGATTGGTGA
GAGCTTAGATCAGAACGATGTATCCTCCTTAGTTTTCCTTACAAGGGATTACACAGGCAGAGGCAAGATAGCCAAGGACA
AGAGTTTCTTGGATCTGGTGATTGAATTGGAGAAACTGAATCTAATTGCTTCAGACCAATTGAATTTGTTAGAAAAATGC
CTGAAGAACATCCACAGAATAGACTTGAACACAAAGATCCAGAAGTACACCCAGTCCAGCCAAGGAGCAAGATCAAATAT
GAATACTCTCCAGGCTTCGCTCCCAAAATTGAGTATCAAGTATAACTCAAGGGTGAGTCTGGAGCCAGTGTATGGAGTAC
CAGCATGA
ORF - retro_mmus_244 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 100. % |
| Parental protein coverage: | 100. % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | MAQSPVSAEVIHQVEECLDEDEKEMMLFLCRDVTENLAAPNVRDLLDSLSERGQLSFATLAELLYRVRRF |
| MAQSPVSAEVIHQVEECLDEDEKEMMLFLCRDVTENLAAPNVRDLLDSLSERGQLSFATLAELLYRVRRF | |
| Retrocopy | MAQSPVSAEVIHQVEECLDEDEKEMMLFLCRDVTENLAAPNVRDLLDSLSERGQLSFATLAELLYRVRRF |
| Parental | DLLKRILKTDKATVEDHLRRNPHLVSDYRVLLMEIGESLDQNDVSSLVFLTRDYTGRGKIAKDKSFLDLV |
| DLLKRILKTDKATVEDHLRRNPHLVSDYRVLLMEIGESLDQNDVSSLVFLTRDYTGRGKIAKDKSFLDLV | |
| Retrocopy | DLLKRILKTDKATVEDHLRRNPHLVSDYRVLLMEIGESLDQNDVSSLVFLTRDYTGRGKIAKDKSFLDLV |
| Parental | IELEKLNLIASDQLNLLEKCLKNIHRIDLNTKIQKYTQSSQGARSNMNTLQASLPKLSIKYNSRVSLEPV |
| IELEKLNLIASDQLNLLEKCLKNIHRIDLNTKIQKYTQSSQGARSNMNTLQASLPKLSIKYNSRVSLEPV | |
| Retrocopy | IELEKLNLIASDQLNLLEKCLKNIHRIDLNTKIQKYTQSSQGARSNMNTLQASLPKLSIKYNSRVSLEPV |
| Parental | YGVPA |
| YGVPA | |
| Retrocopy | YGVPA |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .00 RPM | 18 .85 RPM |
| SRP007412_cerebellum | 0 .00 RPM | 23 .10 RPM |
| SRP007412_heart | 0 .03 RPM | 48 .13 RPM |
| SRP007412_kidney | 0 .08 RPM | 130 .47 RPM |
| SRP007412_liver | 0 .05 RPM | 127 .07 RPM |
| SRP007412_testis | 0 .00 RPM | 10 .84 RPM |
RNA Polymerase II actvity near the 5' end of retro_mmus_244 was not detected
No EST(s) were mapped for retro_mmus_244 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_94690 | 572 libraries | 416 libraries | 77 libraries | 6 libraries | 1 library |
The graphical summary, for retro_mmus_244 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
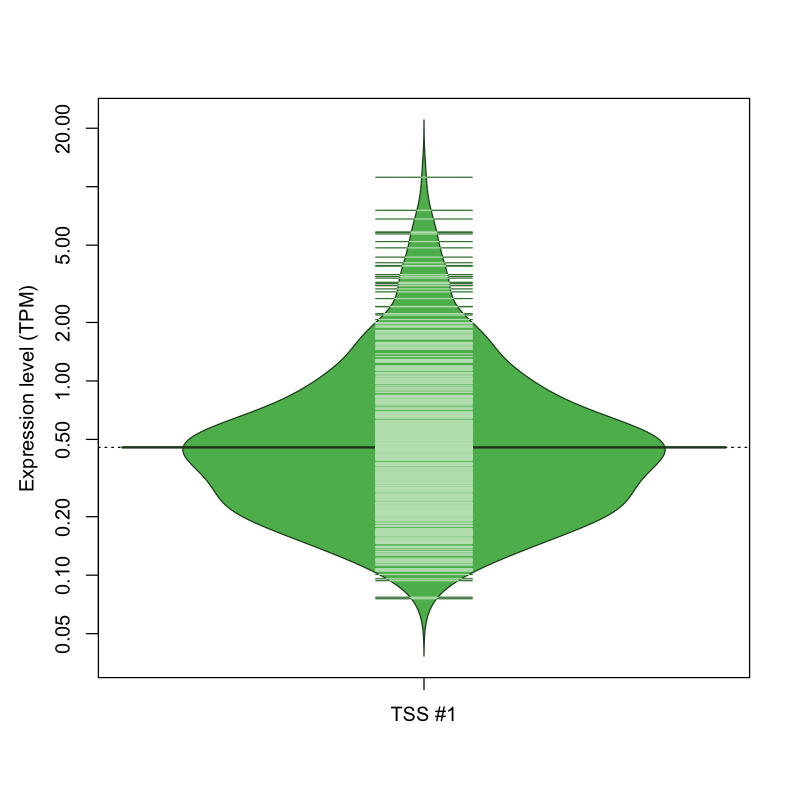
retro_mmus_244 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_244 has 0 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 3 parental genes, and 3 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Callithrix jacchus | ENSCJAG00000004505 | 1 retrocopy | |
| Monodelphis domestica | ENSMODG00000015097 | 1 retrocopy | |
| Mus musculus | ENSMUSG00000026031 | 1 retrocopy |
retro_mmus_244 ,
|