RetrogeneDB ID: | retro_mmus_117 | ||
Retrocopylocation | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 3:124321192..124322284(+) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | ENSMUSG00000044528 | |
| Aliases: | Tram1l1, A830091N21Rik | ||
| Status: | KNOWN_PROTEIN_CODING | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | Tram1 | ||
| Ensembl ID: | ENSMUSG00000025935 | ||
| Aliases: | Tram1, 1810049E02Rik, TRAMP | ||
| Description: | translocating chain-associating membrane protein 1 [Source:MGI Symbol;Acc:MGI:1919515] |

Retrocopy-Parental alignment summary:
>retro_mmus_117
ATGGGGCTCCGCAAGAAGAACGCCAGGAACCCCCCGGTGTTGAGCCACGAATTCATGGTGCAGAACCACGCGGATATGGT
CTCCTGCGTGGGCATGTTCTTCGTGCTGGGGCTTATGTTCGAGGGCACCTCCGAGATGTCGATCGCGTTCCTCACTCTGC
AGCATGGAGTCGTTGTCCCAGCGGAAGGGCTGCCCTCCGGGTCCAGGACCCTGTACCATTATGGGGTCAAAGATCTGGCC
ACAGTGTTCTTCTACATGCTGGTGGCCATCATCATCCACGCCACCATTCAGGAGTACGTGCTAGATAAGCTCAGCAGGAG
ACTGCAGCTCACCAAAGGCAAACAAAACAAACTGAATGAGGCCGGGCAGCTGAGTGTGTTCTATATAGTGTCCGGGATCT
GGGGCATGATCATTCTGGCCTCTGAGAACTGCCTGTCAGACCCTACACTCTTGTGGAAGTCTCAGCCCCACAACATGATG
ACATTTCAGATGAAATTTTTCTACATTTCACAGTTGGCTTACTGGTTTCATAGCTTTCCGGAGCTTTACTTCCAGAAAGT
CAGGAAGCAAGATATCCCGGGTCAGCTCATCTACATTGGCCTCCACCTCTTCCACATTGGAGGGGCCTATCTCTTGTACT
TGAACCACTTGGGCCTGCTGCTGCTGATGCTGCACTACGCCGTAGAGCTCCTCTCCAGCGTGTGCAGCCTGCTTTACTTT
GGTGATGAGCGGTACCAGAAAGGGTTGTCTCTGTGGCCTATCGTGTTTATATCCGGGAGACTCGTGACATTGATTGTCTC
GGTGGTTACAGTAGGGCTTCACCTGGCCGGGACAAATCGGAATGGCAATGCACTCTCTGGTAATGTCAATGTGCTGGCAG
CTAAAATCGCTGTTCTGTCCTCGAGTTGCAGTATCCAGGTGTACATAACATGGACCTTGACTACTGTCTGGCTTCAGAGA
TGGTTAGAAGACGCGAACCTTCATGTTTGTGGAAGGAAGAGACGGTCCAGGGCGAGAAAAGGCACAGAAAATGGAGTGGA
GAATCCGAATAGAATAGACTCTCCACCAAAGAAGAAAGAGAAGGCTCCTTAA
ATGGGGCTCCGCAAGAAGAACGCCAGGAACCCCCCGGTGTTGAGCCACGAATTCATGGTGCAGAACCACGCGGATATGGT
CTCCTGCGTGGGCATGTTCTTCGTGCTGGGGCTTATGTTCGAGGGCACCTCCGAGATGTCGATCGCGTTCCTCACTCTGC
AGCATGGAGTCGTTGTCCCAGCGGAAGGGCTGCCCTCCGGGTCCAGGACCCTGTACCATTATGGGGTCAAAGATCTGGCC
ACAGTGTTCTTCTACATGCTGGTGGCCATCATCATCCACGCCACCATTCAGGAGTACGTGCTAGATAAGCTCAGCAGGAG
ACTGCAGCTCACCAAAGGCAAACAAAACAAACTGAATGAGGCCGGGCAGCTGAGTGTGTTCTATATAGTGTCCGGGATCT
GGGGCATGATCATTCTGGCCTCTGAGAACTGCCTGTCAGACCCTACACTCTTGTGGAAGTCTCAGCCCCACAACATGATG
ACATTTCAGATGAAATTTTTCTACATTTCACAGTTGGCTTACTGGTTTCATAGCTTTCCGGAGCTTTACTTCCAGAAAGT
CAGGAAGCAAGATATCCCGGGTCAGCTCATCTACATTGGCCTCCACCTCTTCCACATTGGAGGGGCCTATCTCTTGTACT
TGAACCACTTGGGCCTGCTGCTGCTGATGCTGCACTACGCCGTAGAGCTCCTCTCCAGCGTGTGCAGCCTGCTTTACTTT
GGTGATGAGCGGTACCAGAAAGGGTTGTCTCTGTGGCCTATCGTGTTTATATCCGGGAGACTCGTGACATTGATTGTCTC
GGTGGTTACAGTAGGGCTTCACCTGGCCGGGACAAATCGGAATGGCAATGCACTCTCTGGTAATGTCAATGTGCTGGCAG
CTAAAATCGCTGTTCTGTCCTCGAGTTGCAGTATCCAGGTGTACATAACATGGACCTTGACTACTGTCTGGCTTCAGAGA
TGGTTAGAAGACGCGAACCTTCATGTTTGTGGAAGGAAGAGACGGTCCAGGGCGAGAAAAGGCACAGAAAATGGAGTGGA
GAATCCGAATAGAATAGACTCTCCACCAAAGAAGAAAGAGAAGGCTCCTTAA
ORF - retro_mmus_117 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 58.52 % |
| Parental protein coverage: | 94.12 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | MAIRKKSNKNPPLLSHEFLLQNHADIVSCLAMLFLLGLMFEVTAKGAIIFVALQYNVTRPATEEQATESA |
| M..RKK...NPP.LSHEF..QNHAD.VSC..M.F.LGLMFE.T....I.F..LQ..V..PA.E.....S. | |
| Retrocopy | MGLRKKNARNPPVLSHEFMVQNHADMVSCVGMFFVLGLMFEGTSEMSIAFLTLQHGVVVPA-EGLPSGSR |
| Parental | SLYHYGIKDLATVLFYMLVAIIIHAIIQEYVLDKINRRMHFSKTKHSKFNESGQLSAFYLFACVWGTFIL |
| .LYHYG.KDLATV.FYMLVAIIIHA.IQEYVLDK..RR....K.K..K.NE.GQLS.FY.....WG..IL | |
| Retrocopy | TLYHYGVKDLATVFFYMLVAIIIHATIQEYVLDKLSRRLQLTKGKQNKLNEAGQLSVFYIVSGIWGMIIL |
| Parental | ISENYISDPTILWRAYPHNLMTFQTKFFYISQLAYWLHAFPELYFQKTKKEDIPRQLVYIGLYLFHIAGA |
| .SEN..SDPT.LW...PHN.MTFQ.KFFYISQLAYW.H.FPELYFQK..K.DIP.QL.YIGL.LFHI.GA | |
| Retrocopy | ASENCLSDPTLLWKSQPHNMMTFQMKFFYISQLAYWFHSFPELYFQKVRKQDIPGQLIYIGLHLFHIGGA |
| Parental | YLLNLNHLGLVLLVLHYFVEFLFHISRLFYFSDEKYQKGFSLWAVLFVLGRLLTLILSVLTVGFGLARAE |
| YLL.LNHLGL.LL.LHY.VE.L.....L.YF.DE.YQKG.SLW...F..GRL.TLI.SV.TVG..LA... | |
| Retrocopy | YLLYLNHLGLLLLMLHYAVELLSSVCSLLYFGDERYQKGLSLWPIVFISGRLVTLIVSVVTVGLHLA-GT |
| Parental | NQKLDFSTGNFNVLAVRIAVLASICITQAFMMWKFINFQLRRWREHSAFQAPPVKRKPAVTKGRSSRKGT |
| N.......GN.NVLA..IAVL.S.C..Q....W......L.RW.E.........KR.....KG.....G. | |
| Retrocopy | NRNGNALSGNVNVLAAKIAVLSSSCSIQVYITWTLTTVWLQRWLEDANLHVCGRKRRSRARKG--TENGV |
| Parental | EN |
| EN | |
| Retrocopy | EN |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 17 .31 RPM | 24 .95 RPM |
| SRP007412_cerebellum | 13 .20 RPM | 24 .32 RPM |
| SRP007412_heart | 0 .16 RPM | 25 .87 RPM |
| SRP007412_kidney | 0 .04 RPM | 69 .93 RPM |
| SRP007412_liver | 0 .00 RPM | 113 .03 RPM |
| SRP007412_testis | 0 .23 RPM | 149 .34 RPM |
RNA Polymerase II actvity may be related with retro_mmus_117 in 1 libraries
| ENCODE library ID | Target | ChIP-Seq Peak coordinates |
|---|---|---|
| ENCFF001XVA | POLR2A | 3:124320791..124321570 |
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_91904 | 529 libraries | 126 libraries | 185 libraries | 80 libraries | 152 libraries |
| TSS #2 | TSS_91905 | 831 libraries | 194 libraries | 47 libraries | 0 libraries | 0 libraries |
The graphical summary, for retro_mmus_117 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
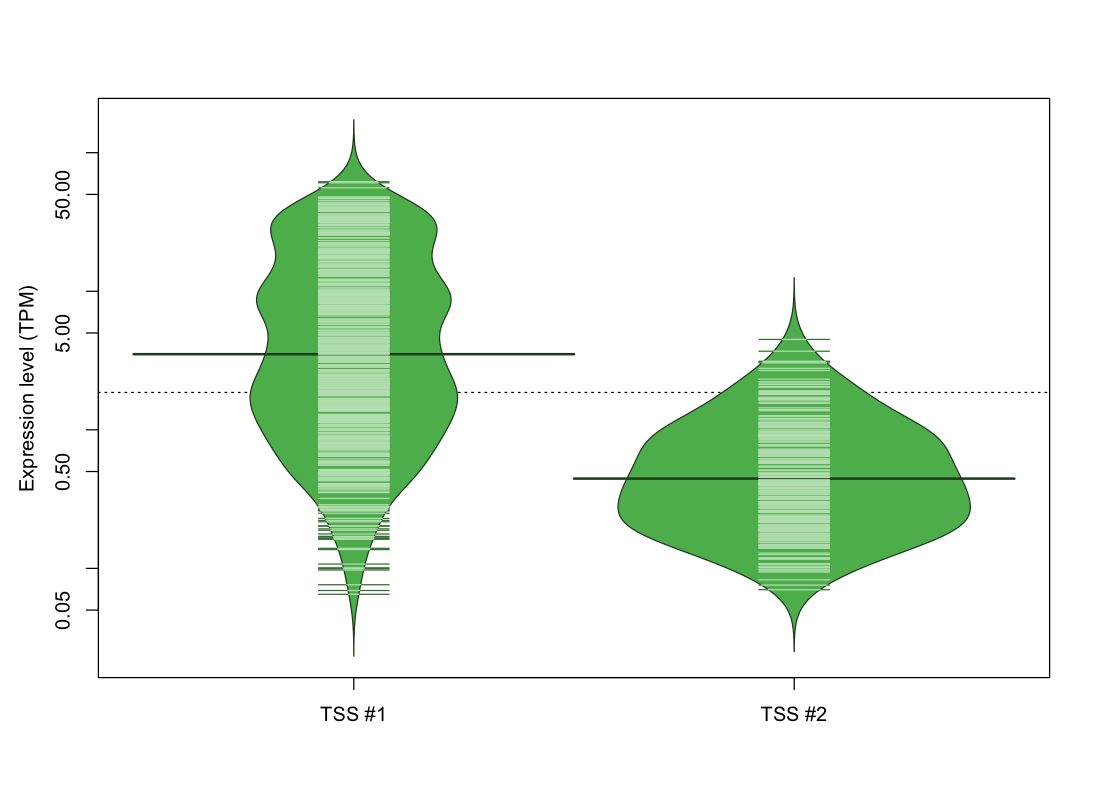
retro_mmus_117 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_117 has 1 orthologous retrocopies within eutheria group .| Species | RetrogeneDB ID |
|---|---|
| Oryctolagus cuniculus | retro_ocun_52 |