RetrogeneDB ID: | retro_hsap_2961 | ||
Retrocopylocation | Organism: | Human (Homo sapiens) | |
| Coordinates: | 4:97823921..97824194(+) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | ENSG00000236764 | |
| Aliases: | None | ||
| Status: | KNOWN_PSEUDOGENE | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | COX7A2 | ||
| Ensembl ID: | ENSG00000112695 | ||
| Aliases: | COX7A2, COX7AL, COX7AL1, COXVIIAL, COXVIIa-L, VIIAL | ||
| Description: | cytochrome c oxidase subunit VIIa polypeptide 2 (liver) [Source:HGNC Symbol;Acc:2288] |

Retrocopy-Parental alignment summary:
>retro_hsap_2961
TGTGGCGGTTGCTGGTCAGTAACAGCCAAGATGCTGTGGAATCTGCTGGCTCTTCATCAGATTGGGCAGAGGACCATAAG
CACTGCTTCCCACAGGCATTTTAAAAATAAAGTTCCAGAGAAACAAAAACTGTTCCAGGAGGATGATGGAATTCCACTGT
ATCTAAAGGGTGGGATAGCTGATGCCCTCCTTCATAGAGCCACCATGATTCTTACAGTTGGTGGAACAGCATATGCCATA
TATCAGCTGGCTGTGGCTTCATTTCCCAATAAA
TGTGGCGGTTGCTGGTCAGTAACAGCCAAGATGCTGTGGAATCTGCTGGCTCTTCATCAGATTGGGCAGAGGACCATAAG
CACTGCTTCCCACAGGCATTTTAAAAATAAAGTTCCAGAGAAACAAAAACTGTTCCAGGAGGATGATGGAATTCCACTGT
ATCTAAAGGGTGGGATAGCTGATGCCCTCCTTCATAGAGCCACCATGATTCTTACAGTTGGTGGAACAGCATATGCCATA
TATCAGCTGGCTGTGGCTTCATTTCCCAATAAA
ORF - retro_hsap_2961 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 91.21 % |
| Parental protein coverage: | 79.13 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | CGGCWSVTAKMLRNLLALRQIGQRTISTASRRHFKNKVPEKQKLFQEDDEIPLYLKGGVADALLYRATMI |
| CGGCWSVTAKML.NLLAL.QIGQRTISTAS.RHFKNKVPEKQKLFQEDD.IPLYLKGG.ADALL.RATMI | |
| Retrocopy | CGGCWSVTAKMLWNLLALHQIGQRTISTASHRHFKNKVPEKQKLFQEDDGIPLYLKGGIADALLHRATMI |
| Parental | LTVGGTAYAIYELAVASFPKK |
| LTVGGTAYAIY.LAVASFP.K | |
| Retrocopy | LTVGGTAYAIYQLAVASFPNK |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| bodymap2_adipose | 0 .00 RPM | 45 .58 RPM |
| bodymap2_adrenal | 0 .00 RPM | 60 .63 RPM |
| bodymap2_brain | 0 .02 RPM | 94 .28 RPM |
| bodymap2_breast | 0 .00 RPM | 70 .80 RPM |
| bodymap2_colon | 0 .00 RPM | 85 .93 RPM |
| bodymap2_heart | 0 .00 RPM | 107 .17 RPM |
| bodymap2_kidney | 0 .00 RPM | 124 .17 RPM |
| bodymap2_liver | 0 .00 RPM | 86 .94 RPM |
| bodymap2_lung | 0 .00 RPM | 73 .40 RPM |
| bodymap2_lymph_node | 0 .00 RPM | 63 .28 RPM |
| bodymap2_ovary | 0 .00 RPM | 44 .18 RPM |
| bodymap2_prostate | 0 .00 RPM | 66 .32 RPM |
| bodymap2_skeletal_muscle | 0 .00 RPM | 25 .79 RPM |
| bodymap2_testis | 0 .00 RPM | 218 .79 RPM |
| bodymap2_thyroid | 0 .00 RPM | 61 .47 RPM |
| bodymap2_white_blood_cells | 0 .00 RPM | 51 .68 RPM |
RNA Polymerase II actvity near the 5' end of retro_hsap_2961 was not detected
No EST(s) were mapped for retro_hsap_2961 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_147109 | 621 libraries | 672 libraries | 470 libraries | 50 libraries | 16 libraries |
The graphical summary, for retro_hsap_2961 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1829 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
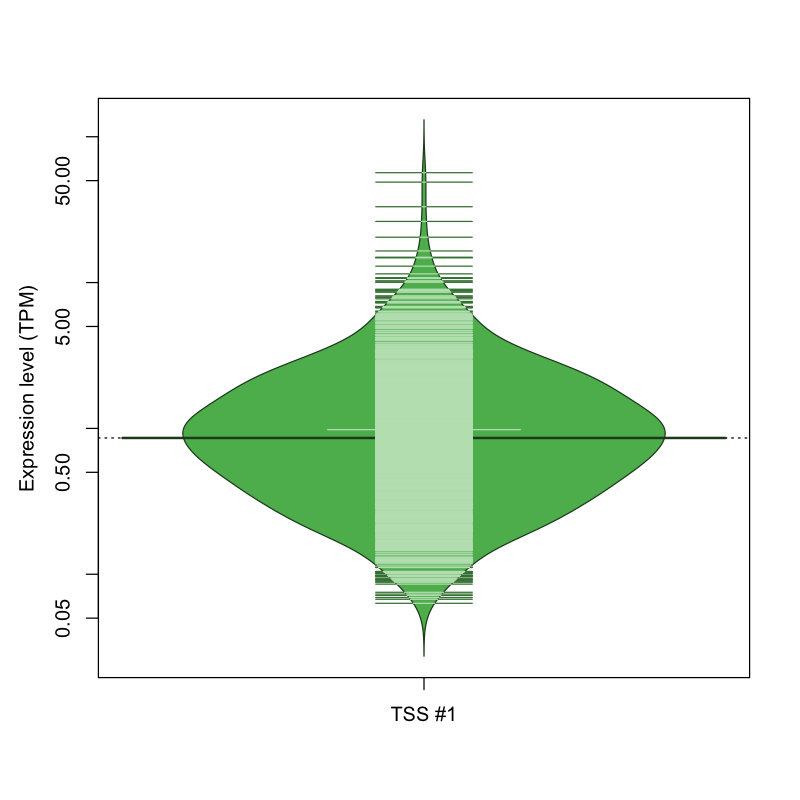
retro_hsap_2961 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_hsap_2961 has 3 orthologous retrocopies within eutheria group .| Species | RetrogeneDB ID |
|---|---|
| Pan troglodytes | retro_ptro_125 |
| Gorilla gorilla | retro_ggor_2051 |
| Pongo abelii | retro_pabe_2456 |
Parental genes homology:
Parental genes homology involve 18 parental genes, and 47 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Ailuropoda melanoleuca | ENSAMEG00000014453 | 1 retrocopy | |
| Bos taurus | ENSBTAG00000005096 | 6 retrocopies | |
| Callithrix jacchus | ENSCJAG00000010305 | 2 retrocopies | |
| Cavia porcellus | ENSCPOG00000007404 | 2 retrocopies | |
| Dipodomys ordii | ENSDORG00000015425 | 2 retrocopies | |
| Homo sapiens | ENSG00000112695 | 2 retrocopies |
retro_hsap_1434, retro_hsap_2961 ,
|
| Homo sapiens | ENSG00000115944 | 1 retrocopy | |
| Gorilla gorilla | ENSGGOG00000005865 | 2 retrocopies | |
| Loxodonta africana | ENSLAFG00000002313 | 1 retrocopy | |
| Microcebus murinus | ENSMICG00000015747 | 3 retrocopies | |
| Nomascus leucogenys | ENSNLEG00000004949 | 2 retrocopies | |
| Pongo abelii | ENSPPYG00000016781 | 2 retrocopies | |
| Pteropus vampyrus | ENSPVAG00000010401 | 6 retrocopies | |
| Rattus norvegicus | ENSRNOG00000042903 | 1 retrocopy | |
| Sus scrofa | ENSSSCG00000004480 | 1 retrocopy | |
| Ictidomys tridecemlineatus | ENSSTOG00000002536 | 5 retrocopies | |
| Tupaia belangeri | ENSTBEG00000007294 | 7 retrocopies | |
| Tursiops truncatus | ENSTTRG00000007874 | 1 retrocopy |
Expression level across human populations :
| Library | Retrogene expression |
|---|---|
| CEU_NA11831 | 0 .00 RPM |
| CEU_NA11843 | 0 .00 RPM |
| CEU_NA11930 | 0 .00 RPM |
| CEU_NA12004 | 0 .00 RPM |
| CEU_NA12400 | 0 .00 RPM |
| CEU_NA12751 | 0 .00 RPM |
| CEU_NA12760 | 0 .00 RPM |
| CEU_NA12827 | 0 .00 RPM |
| CEU_NA12872 | 0 .00 RPM |
| CEU_NA12873 | 0 .00 RPM |
| FIN_HG00183 | 0 .00 RPM |
| FIN_HG00277 | 0 .00 RPM |
| FIN_HG00315 | 0 .00 RPM |
| FIN_HG00321 | 0 .00 RPM |
| FIN_HG00328 | 0 .00 RPM |
| FIN_HG00338 | 0 .00 RPM |
| FIN_HG00349 | 0 .00 RPM |
| FIN_HG00375 | 0 .00 RPM |
| FIN_HG00377 | 0 .00 RPM |
| FIN_HG00378 | 0 .00 RPM |
| GBR_HG00099 | 0 .00 RPM |
| GBR_HG00111 | 0 .00 RPM |
| GBR_HG00114 | 0 .00 RPM |
| GBR_HG00119 | 0 .00 RPM |
| GBR_HG00131 | 0 .00 RPM |
| GBR_HG00133 | 0 .00 RPM |
| GBR_HG00134 | 0 .00 RPM |
| GBR_HG00137 | 0 .03 RPM |
| GBR_HG00142 | 0 .00 RPM |
| GBR_HG00143 | 0 .00 RPM |
| TSI_NA20512 | 0 .03 RPM |
| TSI_NA20513 | 0 .00 RPM |
| TSI_NA20518 | 0 .00 RPM |
| TSI_NA20532 | 0 .00 RPM |
| TSI_NA20538 | 0 .00 RPM |
| TSI_NA20756 | 0 .00 RPM |
| TSI_NA20765 | 0 .00 RPM |
| TSI_NA20771 | 0 .00 RPM |
| TSI_NA20786 | 0 .00 RPM |
| TSI_NA20798 | 0 .00 RPM |
| YRI_NA18870 | 0 .03 RPM |
| YRI_NA18907 | 0 .00 RPM |
| YRI_NA18916 | 0 .00 RPM |
| YRI_NA19093 | 0 .00 RPM |
| YRI_NA19099 | 0 .03 RPM |
| YRI_NA19114 | 0 .00 RPM |
| YRI_NA19118 | 0 .02 RPM |
| YRI_NA19213 | 0 .00 RPM |
| YRI_NA19214 | 0 .00 RPM |
| YRI_NA19223 | 0 .00 RPM |