RetrogeneDB ID: | retro_hsap_1120 | ||
Retrocopylocation | Organism: | Human (Homo sapiens) | |
| Coordinates: | 12:55805508..55808298(-) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | ENSG00000179899 | |
| Aliases: | None | ||
| Status: | KNOWN_PSEUDOGENE | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | PHC1 | ||
| Ensembl ID: | ENSG00000111752 | ||
| Aliases: | PHC1, EDR1, HPH1, MCPH11, RAE28 | ||
| Description: | polyhomeotic homolog 1 (Drosophila) [Source:HGNC Symbol;Acc:3182] |

Retrocopy-Parental alignment summary:
>retro_hsap_1120
CAGGCCACAATTGCTGCCAGTCGGCAGGCCAGCTCCCCAAACACCAGCACTACACAGCAGCAGACTACCACCACCCAGGC
CTCGATCAATCTGGCCACCACATCGGCCGCCCAGCTCATCAGCCGATCCCAGAGTGTGAGCTCTCCCAGTGCTACCACCT
TGACCCAATCTGTGCTACTGGGGAACACCACCTCCCCACCCCTCAACCAGTCTCAGGCCCAGATGTATCTACGGCCACAG
CTGGGAAACCTATTGCAGGTAAACCGAACCCTGGGTCGGAATGTGCCTCTAGCCTCCCAACTCATCCTGATGCCTAATGG
GGCAGTGGCTGCAGTCCAGCAGGAGGTGCCATCTGCTCAGTCTCCTGGAGTTCATGCAGATGCAGATCAGGTTCAGAACT
TGGCAGTAAGGAATCAACAGGCCTCAGCTCAAGGACCTCAGATGCAAGGCTCCACTCAGAAGGCCATTCCTCCAGGAGCC
TCCCCTGTCTCTAGCCTCTCCCAGGCCTCTAGCCAGGCCCTAGCGGTGGCACAGGCTTCCTCTGGGGCCACAAACCAGTC
CCTCAACCTTAGTCAAGCTGGTGGAGGCAGTGGGAATAGCATCCCAGGGTCCATGGGTCCAGGTGGAGGTGGGCAGGCAC
ATGGTGGTTTGGGTCAGTTGCCTTCCTCAGGAATGGGTGGTGGGAGCTGTCCCAGGAAGGGTACAGGAGTGGTGCAGCCC
TTGCCTGCAGCCCAAACAGTGACTGTGAGCCAGGGCAGCCAGACAGAGGCAGAAAGTGCAGCAGCCAAGAAGGCAGAAGC
AGATGGGAGTGGCCAGCAGAATGTGGGCATGAACCTGACACGGACAGCCACACCTGCGCCCAGCCAGACACTTATTAGCT
CAGCCACCTACACACAGATCCAGCCCCATTCACTGATTCAGCAACAGCAACAGATCCACCTCCAGCAGAAACAGGTGGTG
ATCCAGCAGCAGATTGCCATCCACCACCAGCAGCAGTTCCAGCACTGGCAGTCCCAGCTCCTTCACACAGCTACACACCT
CCAGTTGGCGCAGCAGCAGCAGCAGCAACAACAGCAACAGCAGCAACAGCAGCAGCCGCAAGCCACCACCCTCACTGCCC
CTCAGCCACCACAGGTCCCACCTACTCAGCAGGTCCCACCTTCCCAGTCCCAGCAGCAAGCCCAAACCCTGGTCGTTCAG
CCCATGCTTCAGTCTTCACCCTTGTCTCTTCCACCTGATGCAGCCCCTAAGCCACCAATTCCCATCCAATCCAAACCACC
TGTAGCACCTATCAAGCCGCCTCAGTTAGGGGCCGCTAAGATGTCAGCTGCCCAGCAACCACCACCCCATATCCCTGTGC
AAGTTGTAGGCACTCGACAGCCAGGTACAGCCCAGGCACAGGCTTTGGGGTTGGCACAGCTGGCAGCTGCTGTACCTACT
TCCCGGGGGATGCCAGGTACAGTGCAGTCTGGTCAGGCCCATTTGGCCTCCTCGCCACCTTCATCCCAGGCTCCTGGTGC
ACTGCAGGAGTGCCCTCCCACATTGGCCCCTGGGATGACCCTTGCTCCTGTGCAGGGGACAGCACATGTGGTAAAGGGTG
GGGCTACCACCTCCTCACCTGTTGTAGCCCAGGTCCCTGCTGCCTTCTATATGCAGTCTGTGCACTTGCCGGGTAAACCC
CAGACATTGGCTGTCAAACGCAAGGCTGACTCTGAGGAGGAGAGAGATGATGTCTCCACATTGGGTTCAATGCTTCCTGC
CAAGGCATCTCCAGTAGCAGAAAGCCCAAAAGTCATGGACGAGAAGAGCAGTCTTGGAGAAAAAGCTGAATCAGTGGCTA
ATGTGAATGCTAATACTCCAAACAGTGAACTAGTAGCCTTGACCCCCGCCCCTTCAGTACCGCCTCCTACACTAGCCATG
GTGTCTAGACAAATGGGTGACTCAAAACCCCCACAGGCCATCGTGAAGCCCCAGATTCTCACCCACATCATTGAAGGCTT
TGTTATCCAGGAAGGAGCAGAACCTTTCCCGGTGGGTTGTTCTCAGTTACTGAAGGAGTCTGAGAAGCCACTACAGACTG
GCCTTCCGACAGGGCTGACTGAGAATCAGTCAGGTGGCCCTTTGGGAGTGGACAGCCCATCTGCTGAGTTAGATAAGAAG
GCGAATCTCCTGAAGTGCGAGTACTGTGGGAAGTACGCCCCCGCAGAGCAGTTTCGTGGCTCTAAGAGGTTCTGCTCCAT
GACTTGCGCTAAGAGGTACAATGTGAGCTGTAGCCATCAGTTCCGGCTGAAGAGGAAAAAAATGAAAGAGTTTCAAGAAG
CCAACTATGCTCGCGTTCGCAGGCGTGGACCCCGCCGCAGCTCCTCTGACATTGCCCGTGCCAAGATTCAGGGCAAGTGC
CACCGGGGTCAAGAAGACTCTAGCCGGGGTTCAGATAATTCCAGTTATGATGAAGCACTCTCTCCAACATCTCCTGGGCC
TTTATCAGTAAGAGCTGGGCATGGAGAACGTGACCTGGGGAATCCCAATACAGCTCCACCTACACCGGAATTACATGGCA
TCAACCCTGTGTTCCTGTCCAGTAATCCCAGCCGTTGGAGTGTAGAGGAGGTGTACGAGTTTATTGCTTCTCTCCAAGGC
TGCCAAGAGATTGCAGAGGAATTTCGCTCACAGGAGATTGATGGACAGGCCCTTTTATTACTTAAAGAAGAACATCTTAT
GAGTGCCATGAACATCAAGCTGGGCCCTGCCCTCAAGATCTGCGCCAAGATAAATGTCCTCAAGGAGACC
CAGGCCACAATTGCTGCCAGTCGGCAGGCCAGCTCCCCAAACACCAGCACTACACAGCAGCAGACTACCACCACCCAGGC
CTCGATCAATCTGGCCACCACATCGGCCGCCCAGCTCATCAGCCGATCCCAGAGTGTGAGCTCTCCCAGTGCTACCACCT
TGACCCAATCTGTGCTACTGGGGAACACCACCTCCCCACCCCTCAACCAGTCTCAGGCCCAGATGTATCTACGGCCACAG
CTGGGAAACCTATTGCAGGTAAACCGAACCCTGGGTCGGAATGTGCCTCTAGCCTCCCAACTCATCCTGATGCCTAATGG
GGCAGTGGCTGCAGTCCAGCAGGAGGTGCCATCTGCTCAGTCTCCTGGAGTTCATGCAGATGCAGATCAGGTTCAGAACT
TGGCAGTAAGGAATCAACAGGCCTCAGCTCAAGGACCTCAGATGCAAGGCTCCACTCAGAAGGCCATTCCTCCAGGAGCC
TCCCCTGTCTCTAGCCTCTCCCAGGCCTCTAGCCAGGCCCTAGCGGTGGCACAGGCTTCCTCTGGGGCCACAAACCAGTC
CCTCAACCTTAGTCAAGCTGGTGGAGGCAGTGGGAATAGCATCCCAGGGTCCATGGGTCCAGGTGGAGGTGGGCAGGCAC
ATGGTGGTTTGGGTCAGTTGCCTTCCTCAGGAATGGGTGGTGGGAGCTGTCCCAGGAAGGGTACAGGAGTGGTGCAGCCC
TTGCCTGCAGCCCAAACAGTGACTGTGAGCCAGGGCAGCCAGACAGAGGCAGAAAGTGCAGCAGCCAAGAAGGCAGAAGC
AGATGGGAGTGGCCAGCAGAATGTGGGCATGAACCTGACACGGACAGCCACACCTGCGCCCAGCCAGACACTTATTAGCT
CAGCCACCTACACACAGATCCAGCCCCATTCACTGATTCAGCAACAGCAACAGATCCACCTCCAGCAGAAACAGGTGGTG
ATCCAGCAGCAGATTGCCATCCACCACCAGCAGCAGTTCCAGCACTGGCAGTCCCAGCTCCTTCACACAGCTACACACCT
CCAGTTGGCGCAGCAGCAGCAGCAGCAACAACAGCAACAGCAGCAACAGCAGCAGCCGCAAGCCACCACCCTCACTGCCC
CTCAGCCACCACAGGTCCCACCTACTCAGCAGGTCCCACCTTCCCAGTCCCAGCAGCAAGCCCAAACCCTGGTCGTTCAG
CCCATGCTTCAGTCTTCACCCTTGTCTCTTCCACCTGATGCAGCCCCTAAGCCACCAATTCCCATCCAATCCAAACCACC
TGTAGCACCTATCAAGCCGCCTCAGTTAGGGGCCGCTAAGATGTCAGCTGCCCAGCAACCACCACCCCATATCCCTGTGC
AAGTTGTAGGCACTCGACAGCCAGGTACAGCCCAGGCACAGGCTTTGGGGTTGGCACAGCTGGCAGCTGCTGTACCTACT
TCCCGGGGGATGCCAGGTACAGTGCAGTCTGGTCAGGCCCATTTGGCCTCCTCGCCACCTTCATCCCAGGCTCCTGGTGC
ACTGCAGGAGTGCCCTCCCACATTGGCCCCTGGGATGACCCTTGCTCCTGTGCAGGGGACAGCACATGTGGTAAAGGGTG
GGGCTACCACCTCCTCACCTGTTGTAGCCCAGGTCCCTGCTGCCTTCTATATGCAGTCTGTGCACTTGCCGGGTAAACCC
CAGACATTGGCTGTCAAACGCAAGGCTGACTCTGAGGAGGAGAGAGATGATGTCTCCACATTGGGTTCAATGCTTCCTGC
CAAGGCATCTCCAGTAGCAGAAAGCCCAAAAGTCATGGACGAGAAGAGCAGTCTTGGAGAAAAAGCTGAATCAGTGGCTA
ATGTGAATGCTAATACTCCAAACAGTGAACTAGTAGCCTTGACCCCCGCCCCTTCAGTACCGCCTCCTACACTAGCCATG
GTGTCTAGACAAATGGGTGACTCAAAACCCCCACAGGCCATCGTGAAGCCCCAGATTCTCACCCACATCATTGAAGGCTT
TGTTATCCAGGAAGGAGCAGAACCTTTCCCGGTGGGTTGTTCTCAGTTACTGAAGGAGTCTGAGAAGCCACTACAGACTG
GCCTTCCGACAGGGCTGACTGAGAATCAGTCAGGTGGCCCTTTGGGAGTGGACAGCCCATCTGCTGAGTTAGATAAGAAG
GCGAATCTCCTGAAGTGCGAGTACTGTGGGAAGTACGCCCCCGCAGAGCAGTTTCGTGGCTCTAAGAGGTTCTGCTCCAT
GACTTGCGCTAAGAGGTACAATGTGAGCTGTAGCCATCAGTTCCGGCTGAAGAGGAAAAAAATGAAAGAGTTTCAAGAAG
CCAACTATGCTCGCGTTCGCAGGCGTGGACCCCGCCGCAGCTCCTCTGACATTGCCCGTGCCAAGATTCAGGGCAAGTGC
CACCGGGGTCAAGAAGACTCTAGCCGGGGTTCAGATAATTCCAGTTATGATGAAGCACTCTCTCCAACATCTCCTGGGCC
TTTATCAGTAAGAGCTGGGCATGGAGAACGTGACCTGGGGAATCCCAATACAGCTCCACCTACACCGGAATTACATGGCA
TCAACCCTGTGTTCCTGTCCAGTAATCCCAGCCGTTGGAGTGTAGAGGAGGTGTACGAGTTTATTGCTTCTCTCCAAGGC
TGCCAAGAGATTGCAGAGGAATTTCGCTCACAGGAGATTGATGGACAGGCCCTTTTATTACTTAAAGAAGAACATCTTAT
GAGTGCCATGAACATCAAGCTGGGCCCTGCCCTCAAGATCTGCGCCAAGATAAATGTCCTCAAGGAGACC
ORF - retro_hsap_1120 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 97.85 % |
| Parental protein coverage: | 92.63 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | QATIAASRQASSPNTSTTQQQTTTTQASINLATTSAAQLISRSQSVSSPSATTLTQSVLLGNTTSPPLNQ |
| QATIAASRQASSPNTSTTQQQTTTTQASINLATTSAAQLISRSQSVSSPSATTLTQSVLLGNTTSPPLNQ | |
| Retrocopy | QATIAASRQASSPNTSTTQQQTTTTQASINLATTSAAQLISRSQSVSSPSATTLTQSVLLGNTTSPPLNQ |
| Parental | SQAQMYLRPQLGNLLQVNRTLGRNVPLASQLILMPNGAVAAVQQEVPSAQSPGVHADADQVQNLAVRNQQ |
| SQAQMYLRPQLGNLLQVNRTLGRNVPLASQLILMPNGAVAAVQQEVPSAQSPGVHADADQVQNLAVRNQQ | |
| Retrocopy | SQAQMYLRPQLGNLLQVNRTLGRNVPLASQLILMPNGAVAAVQQEVPSAQSPGVHADADQVQNLAVRNQQ |
| Parental | ASAQGPQMQGSTQKAIPPGASPVSSLSQASSQALAVAQASSGATNQSLNLSQAGGGSGNSIPGSMGPGGG |
| ASAQGPQMQGSTQKAIPPGASPVSSLSQASSQALAVAQASSGATNQSLNLSQAGGGSGNSIPGSMGPGGG | |
| Retrocopy | ASAQGPQMQGSTQKAIPPGASPVSSLSQASSQALAVAQASSGATNQSLNLSQAGGGSGNSIPGSMGPGGG |
| Parental | GQAHGGLGQLPSSGMGGGSCPRKGTGVVQPLPAAQTVTVSQGSQTEAESAAAKKAEADGSGQQNVGMNLT |
| GQAHGGLGQLPSSGMGGGSCPRKGTGVVQPLPAAQTVTVSQGSQTEAESAAAKKAEADGSGQQNVGMNLT | |
| Retrocopy | GQAHGGLGQLPSSGMGGGSCPRKGTGVVQPLPAAQTVTVSQGSQTEAESAAAKKAEADGSGQQNVGMNLT |
| Parental | RTATPAPSQTLISSATYTQIQPHSLIQQQQQIHLQQKQVVIQQQIAIHHQQQFQHRQSQLLHTATHLQLA |
| RTATPAPSQTLISSATYTQIQPHSLIQQQQQIHLQQKQVVIQQQIAIHHQQQFQH.QSQLLHTATHLQL. | |
| Retrocopy | RTATPAPSQTLISSATYTQIQPHSLIQQQQQIHLQQKQVVIQQQIAIHHQQQFQHWQSQLLHTATHLQLX |
| Parental | QQQQQQQQQQQQQQQPQATTLTAPQPPQVPPTQQVPPSQSQQQAQTLVVQPMLQSSPLSLPPDAAPKPPI |
| .................ATTLTAPQPPQVPPTQQVPPSQSQQQAQTLVVQPMLQSSPLSLPPDAAPKPPI | |
| Retrocopy | XXXXXXXXXXXXXXXXXATTLTAPQPPQVPPTQQVPPSQSQQQAQTLVVQPMLQSSPLSLPPDAAPKPPI |
| Parental | PIQSKPPVAPIKPPQLGAAKMSAAQQPPPHIPVQVVGTRQPGTAQAQALGLAQLAAAVPTSRGMPGTVQS |
| PIQSKPPVAPIKPPQLGAAKMSAAQQPPPHIPVQVVGTRQPGTAQAQALGLAQLAAAVPTSRGMPGTVQS | |
| Retrocopy | PIQSKPPVAPIKPPQLGAAKMSAAQQPPPHIPVQVVGTRQPGTAQAQALGLAQLAAAVPTSRGMPGTVQS |
| Parental | GQAHLASSPPSSQAPGALQECPPTLAPGMTLAPVQGTAHVVKGGATTSSPVVAQVPAAFYMQSVHLPGKP |
| GQAHLASSPPSSQAPGALQECPPTLAPGMTLAPVQGTAHVVKGGATTSSPVVAQVPAAFYMQSVHLPGKP | |
| Retrocopy | GQAHLASSPPSSQAPGALQECPPTLAPGMTLAPVQGTAHVVKGGATTSSPVVAQVPAAFYMQSVHLPGKP |
| Parental | QTLAVKRKADSEEERDDVSTLGSMLPAKASPVAESPKVMDEKSSLGEKAESVANVNANTPSSELVALTPA |
| QTLAVKRKADSEEERDDVSTLGSMLPAKASPVAESPKVMDEKSSLGEKAESVANVNANTP.SELVALTPA | |
| Retrocopy | QTLAVKRKADSEEERDDVSTLGSMLPAKASPVAESPKVMDEKSSLGEKAESVANVNANTPNSELVALTPA |
| Parental | PSVPPPTLAMVSRQMGDSKPPQAIVKPQILTHIIEGFVIQEGAEPFPVGCSQLLKESEKPLQTGLPTGLT |
| PSVPPPTLAMVSRQMGDSKPPQAIVKPQILTHIIEGFVIQEGAEPFPVGCSQLLKESEKPLQTGLPTGLT | |
| Retrocopy | PSVPPPTLAMVSRQMGDSKPPQAIVKPQILTHIIEGFVIQEGAEPFPVGCSQLLKESEKPLQTGLPTGLT |
| Parental | ENQSGGPLGVDSPSAELDKKANLLKCEYCGKYAPAEQFRGSKRFCSMTCAKRYNVSCSHQFRLKRKKMKE |
| ENQSGGPLGVDSPSAELDKKANLLKCEYCGKYAPAEQFRGSKRFCSMTCAKRYNVSCSHQFRLKRKKMKE | |
| Retrocopy | ENQSGGPLGVDSPSAELDKKANLLKCEYCGKYAPAEQFRGSKRFCSMTCAKRYNVSCSHQFRLKRKKMKE |
| Parental | FQEANYARVRRRGPRRSSSDIARAKIQGKCHRGQEDSSRGSDNSSYDEALSPTSPGPLSVRAGHGERDLG |
| FQEANYARVRRRGPRRSSSDIARAKIQGKCHRGQEDSSRGSDNSSYDEALSPTSPGPLSVRAGHGERDLG | |
| Retrocopy | FQEANYARVRRRGPRRSSSDIARAKIQGKCHRGQEDSSRGSDNSSYDEALSPTSPGPLSVRAGHGERDLG |
| Parental | NPNTAPPTPELHGINPVFLSSNPSRWSVEEVYEFIASLQGCQEIAEEFRSQEIDGQALLLLKEEHLMSAM |
| NPNTAPPTPELHGINPVFLSSNPSRWSVEEVYEFIASLQGCQEIAEEFRSQEIDGQALLLLKEEHLMSAM | |
| Retrocopy | NPNTAPPTPELHGINPVFLSSNPSRWSVEEVYEFIASLQGCQEIAEEFRSQEIDGQALLLLKEEHLMSAM |
| Parental | NIKLGPALKICAKINVLKET |
| NIKLGPALKICAKINVLKET | |
| Retrocopy | NIKLGPALKICAKINVLKET |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| bodymap2_adipose | 1 .05 RPM | 6 .78 RPM |
| bodymap2_adrenal | 0 .96 RPM | 8 .27 RPM |
| bodymap2_brain | 0 .91 RPM | 5 .86 RPM |
| bodymap2_breast | 0 .47 RPM | 5 .84 RPM |
| bodymap2_colon | 0 .81 RPM | 8 .97 RPM |
| bodymap2_heart | 0 .50 RPM | 7 .46 RPM |
| bodymap2_kidney | 0 .17 RPM | 8 .33 RPM |
| bodymap2_liver | 0 .00 RPM | 0 .80 RPM |
| bodymap2_lung | 0 .25 RPM | 8 .56 RPM |
| bodymap2_lymph_node | 0 .57 RPM | 15 .31 RPM |
| bodymap2_ovary | 1 .48 RPM | 18 .37 RPM |
| bodymap2_prostate | 0 .74 RPM | 12 .28 RPM |
| bodymap2_skeletal_muscle | 0 .29 RPM | 7 .71 RPM |
| bodymap2_testis | 1 .56 RPM | 27 .90 RPM |
| bodymap2_thyroid | 1 .51 RPM | 9 .21 RPM |
| bodymap2_white_blood_cells | 0 .12 RPM | 5 .91 RPM |
RNA Polymerase II actvity near the 5' end of retro_hsap_1120 was not detected
18 EST(s) were mapped to retro_hsap_1120 retrocopy
| EST ID | Start | End | Identity | Match | Mis-match | Score |
|---|---|---|---|---|---|---|
| AA300989 | 55805664 | 55805931 | 99.3 | 262 | 1 | 259 |
| AA338510 | 55805712 | 55806002 | 100 | 289 | 0 | 289 |
| BF759538 | 55807395 | 55807810 | 97.9 | 406 | 7 | 395 |
| BF922901 | 55807563 | 55807808 | 96.7 | 236 | 2 | 228 |
| BI009099 | 55807568 | 55807906 | 98.9 | 334 | 4 | 330 |
| BI752280 | 55805398 | 55806223 | 99.9 | 822 | 1 | 819 |
| BP241296 | 55805426 | 55805978 | 99.5 | 549 | 3 | 546 |
| BQ326716 | 55807411 | 55807737 | 95.7 | 320 | 2 | 307 |
| CN388294 | 55805645 | 55806169 | 100 | 524 | 0 | 524 |
| CN388295 | 55805657 | 55806138 | 100 | 481 | 0 | 481 |
| CN388300 | 55805890 | 55806420 | 100 | 530 | 0 | 530 |
| CN388308 | 55805877 | 55806421 | 100 | 544 | 0 | 544 |
| CT001603 | 55805594 | 55806282 | 100 | 680 | 0 | 680 |
| CX760667 | 55806447 | 55806566 | 99.2 | 117 | 1 | 115 |
| DA609879 | 55805739 | 55806294 | 100 | 555 | 0 | 555 |
| HY270459 | 55805801 | 55806274 | 100 | 473 | 0 | 473 |
| HY306555 | 55805479 | 55806004 | 100 | 523 | 0 | 523 |
| HY310658 | 55806744 | 55807255 | 100 | 511 | 0 | 511 |
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_26866 | 1406 libraries | 363 libraries | 53 libraries | 7 libraries | 0 libraries |
The graphical summary, for retro_hsap_1120 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1829 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
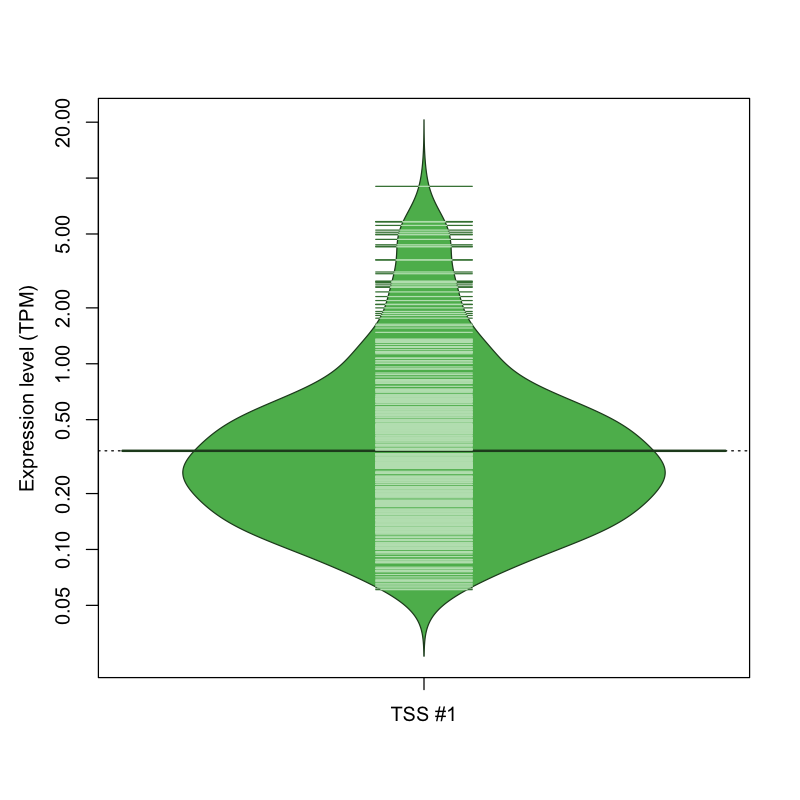
retro_hsap_1120 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_hsap_1120 has 0 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 1 parental gene, and 1 retrocopy.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Homo sapiens | ENSG00000111752 | 1 retrocopy |
retro_hsap_1120 ,
|
Expression level across human populations :
| Library | Retrogene expression |
|---|---|
| CEU_NA11831 | 0 .36 RPM |
| CEU_NA11843 | 0 .37 RPM |
| CEU_NA11930 | 0 .75 RPM |
| CEU_NA12004 | 0 .19 RPM |
| CEU_NA12400 | 0 .35 RPM |
| CEU_NA12751 | 0 .75 RPM |
| CEU_NA12760 | 0 .76 RPM |
| CEU_NA12827 | 0 .05 RPM |
| CEU_NA12872 | 0 .57 RPM |
| CEU_NA12873 | 0 .61 RPM |
| FIN_HG00183 | 0 .41 RPM |
| FIN_HG00277 | 0 .55 RPM |
| FIN_HG00315 | 0 .33 RPM |
| FIN_HG00321 | 0 .54 RPM |
| FIN_HG00328 | 0 .28 RPM |
| FIN_HG00338 | 0 .49 RPM |
| FIN_HG00349 | 0 .26 RPM |
| FIN_HG00375 | 0 .29 RPM |
| FIN_HG00377 | 0 .28 RPM |
| FIN_HG00378 | 0 .17 RPM |
| GBR_HG00099 | 0 .61 RPM |
| GBR_HG00111 | 0 .21 RPM |
| GBR_HG00114 | 0 .45 RPM |
| GBR_HG00119 | 0 .29 RPM |
| GBR_HG00131 | 0 .66 RPM |
| GBR_HG00133 | 0 .31 RPM |
| GBR_HG00134 | 0 .70 RPM |
| GBR_HG00137 | 0 .38 RPM |
| GBR_HG00142 | 0 .56 RPM |
| GBR_HG00143 | 0 .19 RPM |
| TSI_NA20512 | 0 .20 RPM |
| TSI_NA20513 | 0 .39 RPM |
| TSI_NA20518 | 0 .44 RPM |
| TSI_NA20532 | 0 .27 RPM |
| TSI_NA20538 | 0 .14 RPM |
| TSI_NA20756 | 0 .06 RPM |
| TSI_NA20765 | 0 .46 RPM |
| TSI_NA20771 | 0 .51 RPM |
| TSI_NA20786 | 0 .86 RPM |
| TSI_NA20798 | 0 .28 RPM |
| YRI_NA18870 | 0 .61 RPM |
| YRI_NA18907 | 0 .21 RPM |
| YRI_NA18916 | 0 .29 RPM |
| YRI_NA19093 | 0 .57 RPM |
| YRI_NA19099 | 0 .05 RPM |
| YRI_NA19114 | 0 .84 RPM |
| YRI_NA19118 | 0 .00 RPM |
| YRI_NA19213 | 0 .62 RPM |
| YRI_NA19214 | 0 .37 RPM |
| YRI_NA19223 | 0 .46 RPM |