RetrogeneDB ID: | retro_mmus_762 | ||
Retrocopy location | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 11:16635111..16636066(-) | ||
| Located in intron of: | None | ||
Retrocopy information | Ensembl ID: | ENSMUSG00000048149 | |
| Aliases: | None | ||
| Status: | KNOWN_PSEUDOGENE | ||
Parental gene information | Parental gene summary: | ||
| Parental gene symbol: | Ublcp1 | ||
| Ensembl ID: | ENSMUSG00000041231 | ||
| Aliases: | Ublcp1, 4930527B16Rik, 8430435I17Rik, BC002236 | ||
| Description: | ubiquitin-like domain containing CTD phosphatase 1 [Source:MGI Symbol;Acc:MGI:1933105] |

Retrocopy-Parental alignment summary:
>retro_mmus_762
ATGGCACTCCCCATCATAGTGAAATGGGGTGGGCAGGAGTACTCAGTGACCACGCTCTCAGAAGATGATACTGTGCTGGA
TCTTAAACAGTTCCTCAAGACCCTTACGGGAGTGCTCCCAGAGCGCCAGAAGTTACTAGGGCTCAAAGTTAAAGGCAAAC
CTGCAGAAAATGATGTTAAGCTTGGAGCTCTCAAACTGAAACCAAATACTAAAATCATGATGATGGGAACTCGAGAGGAG
AGCTTGGAAGATGTCTTATGTCCTCCTCCTGACAATGATGATGTCATCAATGATTTTGACATTGAAGATGAAGTAGTTGA
AGTAGAAAATAGGGAGGAAAACCTCCTGAAGGTCTCTCGCAGAGTCAAGGAGTACAAGGTGGAGGTGCTGAACCCTCCCC
GGGAGGGCAAGAAGCTCCTGGTGCTGGATGGTGACTATACACTGTTTGATCATAGGTCTTGTGCAGAGACTGGGGTAGAG
TTAATGAGACCATTATCTTCATGAATTTCTAACATCTGCATATGAAGATTATGACATTGTTATTTGGTCTGCAACAAATA
TGAAGTGGATTGAAGCTAAAATGCAAGAGCTGGGCGTGAGCACAAATGCCAACTATAAGATTACCTTCATGCTGGACAGT
GCGGCCATGATCACTGTGCACACGCCCCGGAGAGGGCTCATTGACGTGAAGCCTCTTGGTGTCATTTGGGGAAAGTTTTC
TGAATTCTACAGCAAAAAGAACACCATTATGTTTGATGATATAGGAAGAAATTTTCTAATGAATCCACAAAATGGACTGA
AGATAAGGCCTTTCATGAAAGCTCATCTAAACCGTGATAAAGACAAGGAACTGGTGAAGCTCACTCAGTACCTCAAGGAG
ATCGCTAAGCTTGATGACTTCTTGGAGCTGAATCACAAGTACTGGGAGAGGTGTCTATCAAAGAAGCAAGGACAG
ORF - retro_mmus_762 Open Reading Frame is not conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 98.75 % |
| Parental protein coverage: | 100.0 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected: | 1 |
Retrocopy - Parental Gene Alignment:
| Parental | MALPIIVKWGGQEYSVTTLSEDDTVLDLKQFLKTLTGVLPERQKLLGLKVKGKPAENDVKLGALKLKPNT |
| MALPIIVKWGGQEYSVTTLSEDDTVLDLKQFLKTLTGVLPERQKLLGLKVKGKPAENDVKLGALKLKPNT | |
| Retrocopy | MALPIIVKWGGQEYSVTTLSEDDTVLDLKQFLKTLTGVLPERQKLLGLKVKGKPAENDVKLGALKLKPNT |
| Parental | KIMMMGTREESLEDVLCPPPDNDDVINDFDIEDEVVEVENREENLLKVSRRVKEYKVEVLNPPREGKKLL |
| KIMMMGTREESLEDVLCPPPDNDDVINDFDIEDEVVEVENREENLLKVSRRVKEYKVEVLNPPREGKKLL | |
| Retrocopy | KIMMMGTREESLEDVLCPPPDNDDVINDFDIEDEVVEVENREENLLKVSRRVKEYKVEVLNPPREGKKLL |
| Parental | VLDVDYTLFDHRSCAETGVELMRP-YLHEFLTSAYEDYDIVIWSATNMKWIEAKMKELGVSTNANYKITF |
| VLD.DYTLFDHRSCAETGVELMRP.YLHEFLTSAYEDYDIVIWSATNMKWIEAKM.ELGVSTNANYKITF | |
| Retrocopy | VLDGDYTLFDHRSCAETGVELMRP>YLHEFLTSAYEDYDIVIWSATNMKWIEAKMQELGVSTNANYKITF |
| Parental | MLDSAAMITVHTPRRGLIDVKPLGVIWGKFSEFYSKKNTIMFDDIGRNFLMNPQNGLKIRPFMKAHLNRD |
| MLDSAAMITVHTPRRGLIDVKPLGVIWGKFSEFYSKKNTIMFDDIGRNFLMNPQNGLKIRPFMKAHLNRD | |
| Retrocopy | MLDSAAMITVHTPRRGLIDVKPLGVIWGKFSEFYSKKNTIMFDDIGRNFLMNPQNGLKIRPFMKAHLNRD |
| Parental | KDKELVKLTQYLKEIAKLDDFLELNHKYWERYLSKKQGQ |
| KDKELVKLTQYLKEIAKLDDFLELNHKYWER.LSKKQGQ | |
| Retrocopy | KDKELVKLTQYLKEIAKLDDFLELNHKYWERCLSKKQGQ |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .35 RPM | 22 .59 RPM |
| SRP007412_cerebellum | 2 .00 RPM | 26 .53 RPM |
| SRP007412_heart | 0 .16 RPM | 14 .04 RPM |
| SRP007412_kidney | 1 .10 RPM | 20 .37 RPM |
| SRP007412_liver | 0 .63 RPM | 13 .44 RPM |
| SRP007412_testis | 1 .14 RPM | 8 .51 RPM |
RNA Polymerase II activity near the 5' end of retro_mmus_762 was not detected
No EST(s) were mapped for retro_mmus_762 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_11207 | 150 libraries | 159 libraries | 640 libraries | 101 libraries | 22 libraries |
The graphical summary, for retro_mmus_762 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
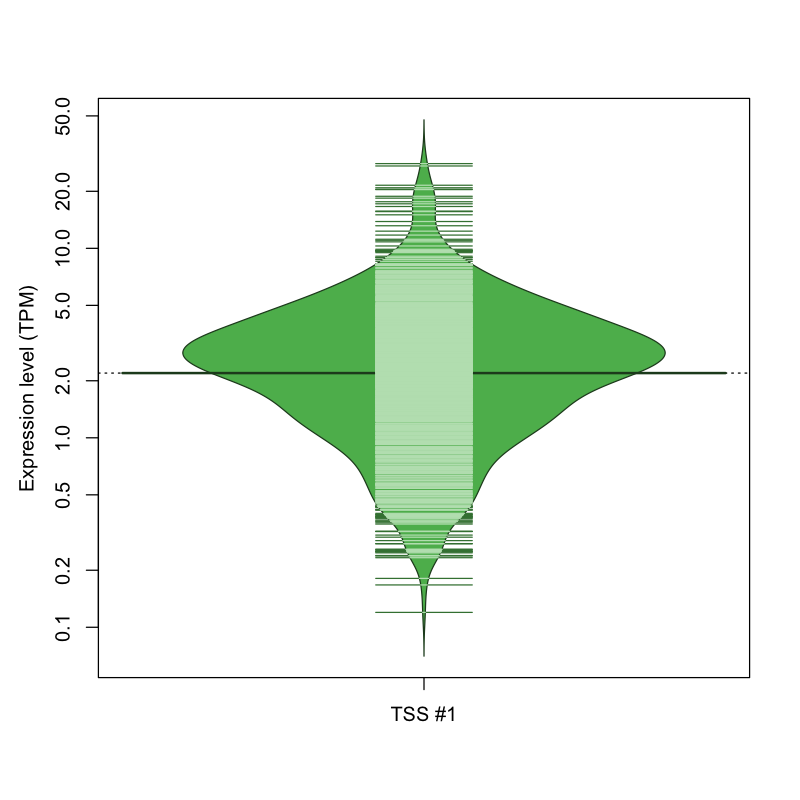
retro_mmus_762 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_762 has 0 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 10 parental genes, and 12 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Choloepus hoffmanni | ENSCHOG00000007691 | 1 retrocopy | |
| Echinops telfairi | ENSETEG00000000170 | 1 retrocopy | |
| Microcebus murinus | ENSMICG00000010060 | 2 retrocopies | |
| Myotis lucifugus | ENSMLUG00000006359 | 1 retrocopy | |
| Monodelphis domestica | ENSMODG00000008063 | 2 retrocopies | |
| Mustela putorius furo | ENSMPUG00000013396 | 1 retrocopy | |
| Mus musculus | ENSMUSG00000041231 | 1 retrocopy |
retro_mmus_762 ,
|
| Oryctolagus cuniculus | ENSOCUG00000002224 | 1 retrocopy | |
| Tupaia belangeri | ENSTBEG00000008983 | 1 retrocopy | |
| Tursiops truncatus | ENSTTRG00000005876 | 1 retrocopy |