RetrogeneDB ID: | retro_mmus_2386 | ||
Retrocopy location | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 4:111290186..111291113(+) | ||
| Located in intron of: | ENSMUSG00000061298 | ||
Retrocopy information | Ensembl ID: | ENSMUSG00000081068 | |
| Aliases: | None | ||
| Status: | KNOWN_PSEUDOGENE | ||
Parental gene information | Parental gene summary: | ||
| Parental gene symbol: | Mfap1a | ||
| Ensembl ID: | ENSMUSG00000068479 | ||
| Aliases: | Mfap1a, 4432409M24Rik, Mfap1 | ||
| Description: | microfibrillar-associated protein 1A [Source:MGI Symbol;Acc:MGI:1914782] |

Retrocopy-Parental alignment summary:
>retro_mmus_2386
GAGAAAGGTGAGATCTCAATGGAAACGGTGAAGGTGAAACGTTATGTGTCTGGAAAGAGACCAGATTATGCCCCTATGGA
GTCCTCAGATGAAGAGGATGAAGAATTTCAGTTCATTAAGAAAGCTAAGGAACAAGAAGCAGAGCCTGAGGAACAGGAGG
AGGATTCATCCAGTGACCCCCGTCTTCGGCGGTTACAGAACCGCATCAGTGAAGATGTGGAAGAGAGAAAGGATCAGATT
ACAGTTCAAGGATGAGAAGCTGAAGCATTGAAACAGAAGGAACTGGAACAAGAAGCAAAATGCGTGGCTGAGGAGTGGCA
CAAGTACACCCTCAAGGTTGTAGAAGAAGAGACCAAGAAAGAGCTAGAGGAGAACAAGCGGTCCCTTGCCGCCCTGGATG
CACTCAATACTGATGATGAAAATGATGAGGAGGAGTACGAGGCGTGGAAAGTCCGCTGGCTCAAAAGAATCAAGTGGGAG
AGAGAAGACTGAGAAGCGCTTGAAAAGGAGAAAGCAGAAATTGAACGCATGAGAAACCTGACTGAGGAAGAAAGGCGGGC
TGAACTTCAGGCAAATGGCAAAGTCATTACCAACAAAGCTGTTAAAGGCAAATACAAGTTTCCACAGAAGTATTATCACC
GAGGTGCCTTCTTCATGGATGAGAATGAAGAAGTCTACAAGAGAGATTTTAGTGCACCTACTCTTGAGGACCATTTCAAC
AAAACCATTCTTCCTAGAGTCATGCAGGTCAAGAATTTTGGACGGTCTGGTCGCACCAAGTACACCCACCTTGTGGATCA
AGATACCACTTCCTTTGACTCTGCATGGGGCCAAGAGAGTGCCCAGAACACAAAATTCTTTAAACAAAAGGCAGCAGGGG
TATGAGATGTATTTGAGTGACCATCTGCCAAGAAGAGGAAAACCACT
ORF - retro_mmus_2386 Open Reading Frame is not conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 74.76 % |
| Parental protein coverage: | 69.25 % |
| Number of stop codons detected: | 4 |
| Number of frameshifts detected: | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | EEEEIDDEEIERRRGMMRQRAQERKNEEMEVMEVEDEG-RSGEESESESEYEEYTDSED----EMEPRLK |
| E..EI..E.....R.....R......E.......E........E.E.E.E..E...S.D........R.. | |
| Retrocopy | EKGEISMETVKVKRYVSGKRPDYAPMESSDEEDEEFQFIKKAKEQEAEPEEQEEDSSSDPRLRRLQNRIS |
| Parental | PVFIRKKDRVTVQEREAEALKQKELEQEAKRMAEERRKYTLKIVEEETKKELEENKRSLAALDALNTDDE |
| ......KD..TVQ..EAEALKQKELEQEAK..AEE..KYTLK.VEEETKKELEENKRSLAALDALNTDDE | |
| Retrocopy | EDVEERKDQITVQG*EAEALKQKELEQEAKCVAEEWHKYTLKVVEEETKKELEENKRSLAALDALNTDDE |
| Parental | NDEEEYEAWKVRELKRIKREREDREALEKEKAEIERMRNLTEEERRAELRANGKVITNKAVKGKYKFLQK |
| NDEEEYEAWKVR.LKRIK.ERED.EALEKEKAEIERMRNLTEEERRAEL.ANGKVITNKAVKGKYKF.QK | |
| Retrocopy | NDEEEYEAWKVRWLKRIKWERED*EALEKEKAEIERMRNLTEEERRAELQANGKVITNKAVKGKYKFPQK |
| Parental | YYHRGAFFMDEDEEVYKRDFSAPTLEDHFNKTILPKVMQVKNFGRSGRTKYTHLVDQDTTSFDSAWGQES |
| YYHRGAFFMDE.EEVYKRDFSAPTLEDHFNKTILP.VMQVKNFGRSGRTKYTHLVDQDTTSFDSAWGQES | |
| Retrocopy | YYHRGAFFMDENEEVYKRDFSAPTLEDHFNKTILPRVMQVKNFGRSGRTKYTHLVDQDTTSFDSAWGQES |
| Parental | AQNTKFFKQKAAGVRDVFERPSAKKRKTT |
| AQNTKFFKQKAAGV.DVFE.PSAKKRKTT | |
| Retrocopy | AQNTKFFKQKAAGV*DVFE*PSAKKRKTT |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .07 RPM | 14 .27 RPM |
| SRP007412_cerebellum | 0 .09 RPM | 17 .80 RPM |
| SRP007412_heart | 0 .09 RPM | 15 .10 RPM |
| SRP007412_kidney | 0 .19 RPM | 14 .90 RPM |
| SRP007412_liver | 0 .00 RPM | 6 .23 RPM |
| SRP007412_testis | 0 .14 RPM | 8 .42 RPM |
RNA Polymerase II activity near the 5' end of retro_mmus_2386 was not detected
No EST(s) were mapped for retro_mmus_2386 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_99177 | 597 libraries | 415 libraries | 56 libraries | 0 libraries | 4 libraries |
The graphical summary, for retro_mmus_2386 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
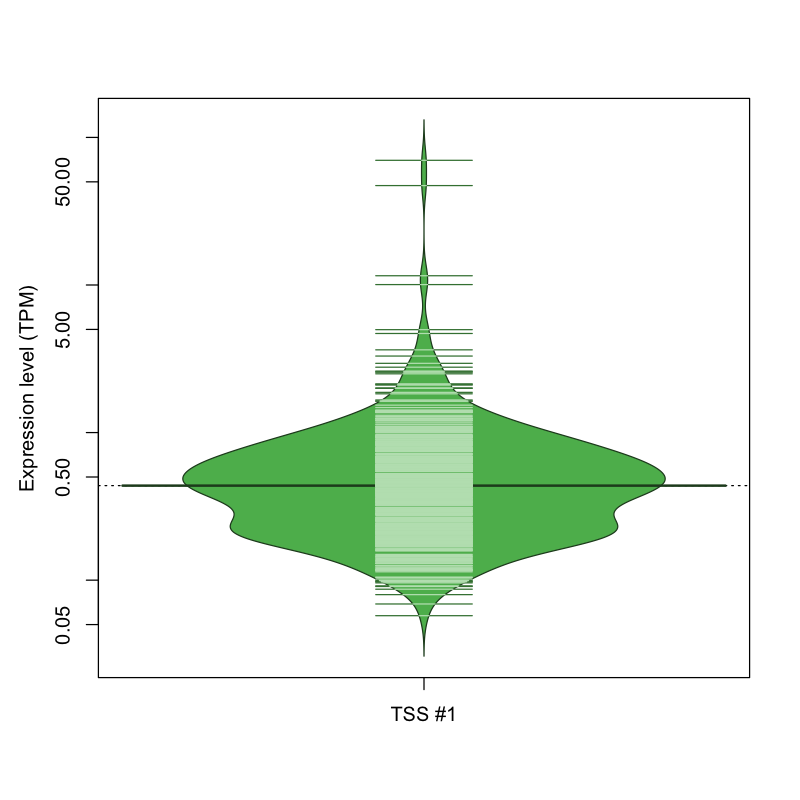
retro_mmus_2386 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_2386 has 0 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 9 parental genes, and 14 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Dipodomys ordii | ENSDORG00000013870 | 1 retrocopy | |
| Myotis lucifugus | ENSMLUG00000013000 | 2 retrocopies | |
| Mus musculus | ENSMUSG00000048222 | 2 retrocopies | |
| Mus musculus | ENSMUSG00000068479 | 2 retrocopies |
retro_mmus_2386 , retro_mmus_2638,
|
| Otolemur garnettii | ENSOGAG00000010206 | 1 retrocopy | |
| Rattus norvegicus | ENSRNOG00000015984 | 1 retrocopy | |
| Sarcophilus harrisii | ENSSHAG00000014628 | 1 retrocopy | |
| Ictidomys tridecemlineatus | ENSSTOG00000006967 | 2 retrocopies | |
| Tupaia belangeri | ENSTBEG00000005050 | 2 retrocopies |