RetrogeneDB ID: | retro_mmus_1604 | ||
Retrocopy location | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 17:3312251..3314068(-) | ||
| Located in intron of: | None | ||
Retrocopy information | Ensembl ID: | None | |
| Aliases: | None | ||
| Status: | NOVEL | ||
Parental gene information | Parental gene summary: | ||
| Parental gene symbol: | Cd2ap | ||
| Ensembl ID: | ENSMUSG00000061665 | ||
| Aliases: | Cd2ap, AL024079, C78928, METS-1, Mets1 | ||
| Description: | CD2-associated protein [Source:MGI Symbol;Acc:MGI:1330281] |

Retrocopy-Parental alignment summary:
>retro_mmus_1604
CTACAGGAAGAAGGATGGCTAGAAGGAGAACTAAATGGGAGATGAGGGATGTTCCCTGACAATTTCGTTAAGGAAATTAA
GCGAGAGACAGAACCCAAGGATGACAATTTGCCCATCAAACGGGAAAGGCGAGGAAATGTAGCAAGTCTTGTACAGCGAA
TCAGTACATATGGACTTCCAGCCGGAGGAATTCAGCCACATCCACAAACCAAAGCTATTAAGAAGAAGACAAAAAAGCGT
CAGTGTAAAGTTCTTTTTGACTACTCTCCACAAAATGAGGACGAACTGGAAATGATCGTGGGGGATGTCATTGACGTCAT
TGAAGAGGTAGAAGAAGGCTGGTGGAGTGGAACCCTGAACAATAAGTCGGGACTGTTTCCCTCAAACTTTGTGAAAGAAT
TAGAGTTCACAGAGGATGGTGAAACTCACAACGCTCAGGAGGAATCAGAAGTTCCTTTGACTGGCCCTACCTCACCGTTG
CCATCTCTGGGGAATGGGAGTGAACCTGCTCCCGGATCAGTTGCACAGCCAAAGAAAATCCGAGGAATTGGATTTGGAGA
CATCTTTAAAGAAGGCTCTGTGAAACTTAGGACAAGAACTTCCAGTAATGAAACAGAGGAGGGGGAAAAAAAAAAGCCTT
TCATCCTACAGTCACTGGGATCTAGAACTCAGAATGTGGAGGTAACAAAACCAGACATTGATGACAAGATTAAAGCTAAG
GAATATTGTAGAACATTACTTCCCTATGCAGAAATCAATGAAGATGAACTTACACTTAGAGAAGGGGAGATAATCCATTT
GATAAGTAAGGAGGCTGGAGAAGCAGGCTGGTGGAAGGGTGAACTGAACGGTAAAGAAGGAGTGTTTCCACATAACTTTG
CTGTTCAGATAAGTGAACTAGATAAAGACTTTCCAAAACCAAAGAAACTACCTCCTCCTGTAAAAGGTCCAGCTCCAAAG
CCTGACCCATAGCTGCAGAGAAGAAAGCTTTTCCTCAAAGGCTGAGGAAAAGATGAAAAATCACTGCTAGAACACAAACC
TTCTAAACCAGCTGCTCCTCAAGTCCCACCCCAAAAGCCCACTCCACCCACCAAAGCCAATAACTTACTGAGGTCGCCTG
GAACAGTGAACCCAAAGCGGCCAGAAAAGCCAGTCCCTCCGCCACCTCCAACAGCCAAAATTAATGGAGAAGTTTCTACA
ATTTCTTCAAAGATTGATACTGAGCCAGTATCAAAACCAAAGCTAGATCCTGAACAACTGCCACTTACACCAAAATCAGT
AGACCTTGATGCCTTTGTAGCCAGGAACTCAAAAGAAACAGATGATGTAAATTTTGATGACATAGCTTCCTCAGAGAACT
TGTTACATCTCACTGCTAATAGACCCAAGATGCCTGGAAGATGACTGCCTGGCCGTTTCAATGGTGGACATTCTCCAACC
CAAAGCCCAGAAAAAACCTTGAAGCTACCAAAAGAAGATGATAGTGGCAACCTAAAGCCTTTGGAGTTTAAAAAGGATGC
AAGCTACTCTTCAAAGCCATCTCTTTCAACACCTTCAAATGCTTCTTATGTAAATACAGCTGCTTTCCTAACTCCCGTTA
GAACTCAAAGCTAAAGCAGAAGCAGATGATGGGGGAAAAAATTCCATGGACAAACTTAGAGCCCAGATTATTGAATTATT
GTGTATTGTAGATGCACTGAAAAAGTATCATGGGGAAAGAACTGGAAAAGCTACGCAAAGAATTAGAAGAAGAGAAGGCC
ATGCGAAGTAATCTAGAGGTGGAAATAGCAAAGCTGAAGAAAGCTGTTCTGTTGTCT
ORF - retro_mmus_1604 Open Reading Frame is not conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 91.49 % |
| Parental protein coverage: | 95.13 % |
| Number of stop codons detected: | 2 |
| Number of frameshifts detected: | 5 |
Retrocopy - Parental Gene Alignment:
| Parental | LQEEGWLEGELNGRRGMFPDNFVKEIKRETEPKDDNLPIKRERQGNVASLVQRISTYGLPAGGIQPHPQT |
| LQEEGWLEGELNGR.GMFPDNFVKEIKRETEPKDDNLPIKRER.GNVASLVQRISTYGLPAGGIQPHPQT | |
| Retrocopy | LQEEGWLEGELNGR*GMFPDNFVKEIKRETEPKDDNLPIKRERRGNVASLVQRISTYGLPAGGIQPHPQT |
| Parental | KAIKKKTKKRQCKVLFDYSPQNEDELELIVGDVIDVIEEVEEGWWSGTLNNKLGLFPSNFVKELESTEDG |
| KAIKKKTKKRQCKVLFDYSPQNEDELE.IVGDVIDVIEEVEEGWWSGTLNNK.GLFPSNFVKELE.TEDG | |
| Retrocopy | KAIKKKTKKRQCKVLFDYSPQNEDELEMIVGDVIDVIEEVEEGWWSGTLNNKSGLFPSNFVKELEFTEDG |
| Parental | ETHNAQEESEVPLTGPTSPLPSPGNGSEPAPGSVAQPKKIRGIGFGDIFKEGSVKLRTRTSSSETEEKKT |
| ETHNAQEESEVPLTGPTSPLPS.GNGSEPAPGSVAQPKKIRGIGFGDIFKEGSVKLRTRTSS.ETEE... | |
| Retrocopy | ETHNAQEESEVPLTGPTSPLPSLGNGSEPAPGSVAQPKKIRGIGFGDIFKEGSVKLRTRTSSNETEEGXX |
| Parental | EKPLILQPLGSRTQNVEVTKPDVDGKIKAKEYCRTLFPYTGTNEDELTFREGEIIHLISKETGEAGWWKG |
| ..P.ILQ.LGSRTQNVEVTKPD.D.KIKAKEYCRTL.PY...NEDELT.REGEIIHLISKE.GEAGWWKG | |
| Retrocopy | XXPFILQSLGSRTQNVEVTKPDIDDKIKAKEYCRTLLPYAEINEDELTLREGEIIHLISKEAGEAGWWKG |
| Parental | ELNGKEGVFPDNFAVQISELDKDFPKPKKPPPPAKGPAPKPD-LSAAEKKAFP-LKAEE-KDEKSLLEQK |
| ELNGKEGVFP.NFAVQISELDKDFPKPKK.PPP.KGPAPKPD...AAEKKAFP..KAEE.KDEKSLLE.K | |
| Retrocopy | ELNGKEGVFPHNFAVQISELDKDFPKPKKLPPPVKGPAPKPD<PIAAEKKAFP<SKAEE<KDEKSLLEHK |
| Parental | PSKPAAPQVPPKKPTAPTKASNLLRSPGAVYPKRPEKPVPPPPPAAKINGEVSIISSKIDTEPVSKPKLD |
| PSKPAAPQVPP.KPT.PTKA.NLLRSPG.V.PKRPEKPVPPPPP.AKINGEVS.ISSKIDTEPVSKPKLD | |
| Retrocopy | PSKPAAPQVPPQKPTPPTKANNLLRSPGTVNPKRPEKPVPPPPPTAKINGEVSTISSKIDTEPVSKPKLD |
| Parental | PEQLPVRPKSVDLDAFVARNSKETDDVNFDDIASSENLLHLTANRPKMPGRRLPGRFNGGHSPTQSPEKT |
| PEQLP..PKSVDLDAFVARNSKETDDVNFDDIASSENLLHLTANRPKMPGR.LPGRFNGGHSPTQSPEKT | |
| Retrocopy | PEQLPLTPKSVDLDAFVARNSKETDDVNFDDIASSENLLHLTANRPKMPGR*LPGRFNGGHSPTQSPEKT |
| Parental | LKLPKEDDSGNLKPLEFKKDASYSSKSSLSTPSSASKVNTAAFLT-PLELKAKAEADDGKRNSVDELRAQ |
| LKLPKEDDSGNLKPLEFKKDASYSSK.SLSTPS.AS.VNTAAFLT.PLELKAKAEADDG..NS.D.LRAQ | |
| Retrocopy | LKLPKEDDSGNLKPLEFKKDASYSSKPSLSTPSNASYVNTAAFLT>PLELKAKAEADDGGKNSMDKLRAQ |
| Parental | IIELLCIVDALKKDH-GKELEKLRKELEEEKAMRSNLEVEIAKLKKAVLLS |
| IIELLCIVDALKK.H.GKELEKLRKELEEEKAMRSNLEVEIAKLKKAVLLS | |
| Retrocopy | IIELLCIVDALKKYH>GKELEKLRKELEEEKAMRSNLEVEIAKLKKAVLLS |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .40 RPM | 23 .95 RPM |
| SRP007412_cerebellum | 0 .30 RPM | 26 .36 RPM |
| SRP007412_heart | 0 .25 RPM | 23 .44 RPM |
| SRP007412_kidney | 0 .33 RPM | 88 .86 RPM |
| SRP007412_liver | 0 .16 RPM | 36 .27 RPM |
| SRP007412_testis | 0 .23 RPM | 18 .34 RPM |
RNA Polymerase II activity may be related with retro_mmus_1604 in 3 libraries
| ENCODE library ID | Target | ChIP-Seq Peak coordinates |
|---|---|---|
| ENCFF001XVA | POLR2A | 17:3314161..3314573 |
| ENCFF001YIJ | POLR2A | 17:3314010..3314958 |
| ENCFF001YKA | POLR2A | 17:3314362..3314570 |
No EST(s) were mapped for retro_mmus_1604 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_52457 | 421 libraries | 481 libraries | 159 libraries | 5 libraries | 6 libraries |
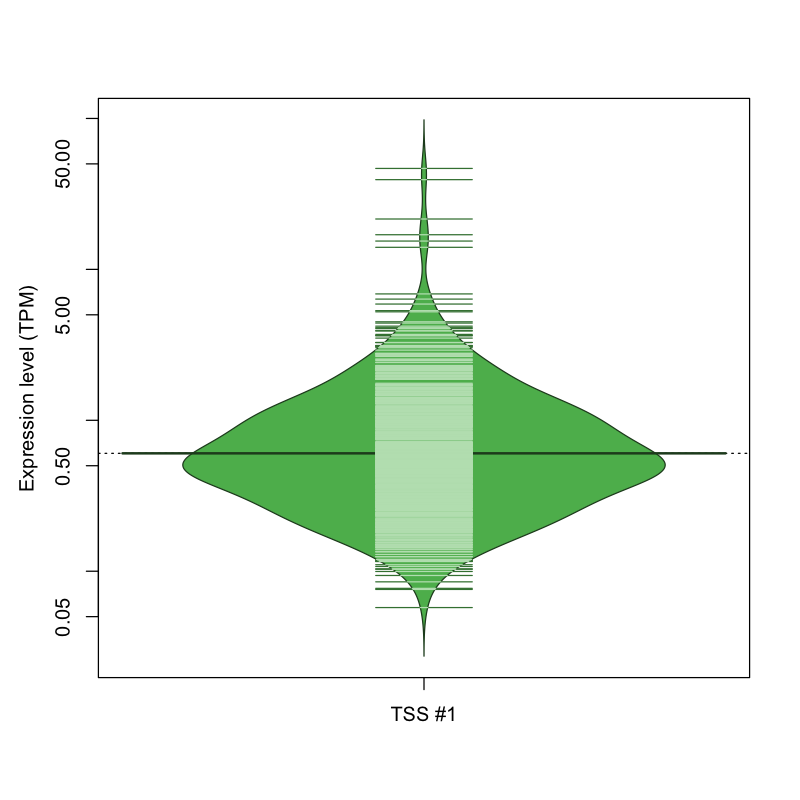
The graphical summary, for retro_mmus_1604 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
retro_mmus_1604 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_1604 has 0 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 4 parental genes, and 4 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Myotis lucifugus | ENSMLUG00000009776 | 1 retrocopy | |
| Mus musculus | ENSMUSG00000061665 | 1 retrocopy |
retro_mmus_1604 ,
|
| Oryctolagus cuniculus | ENSOCUG00000005026 | 1 retrocopy | |
| Rattus norvegicus | ENSRNOG00000011987 | 1 retrocopy |